Mohd Faheem Khan, Mohd Tasleem Khan
Molecules· 2026-01· University College Dublin
The research
Conventional enzyme engineering, whether through directed evolution or rational design, relies on iterative cycles of random mutagenesis and high-throughput screening that are labor-intensive, expensive, and confined to narrow regions of protein sequence space.
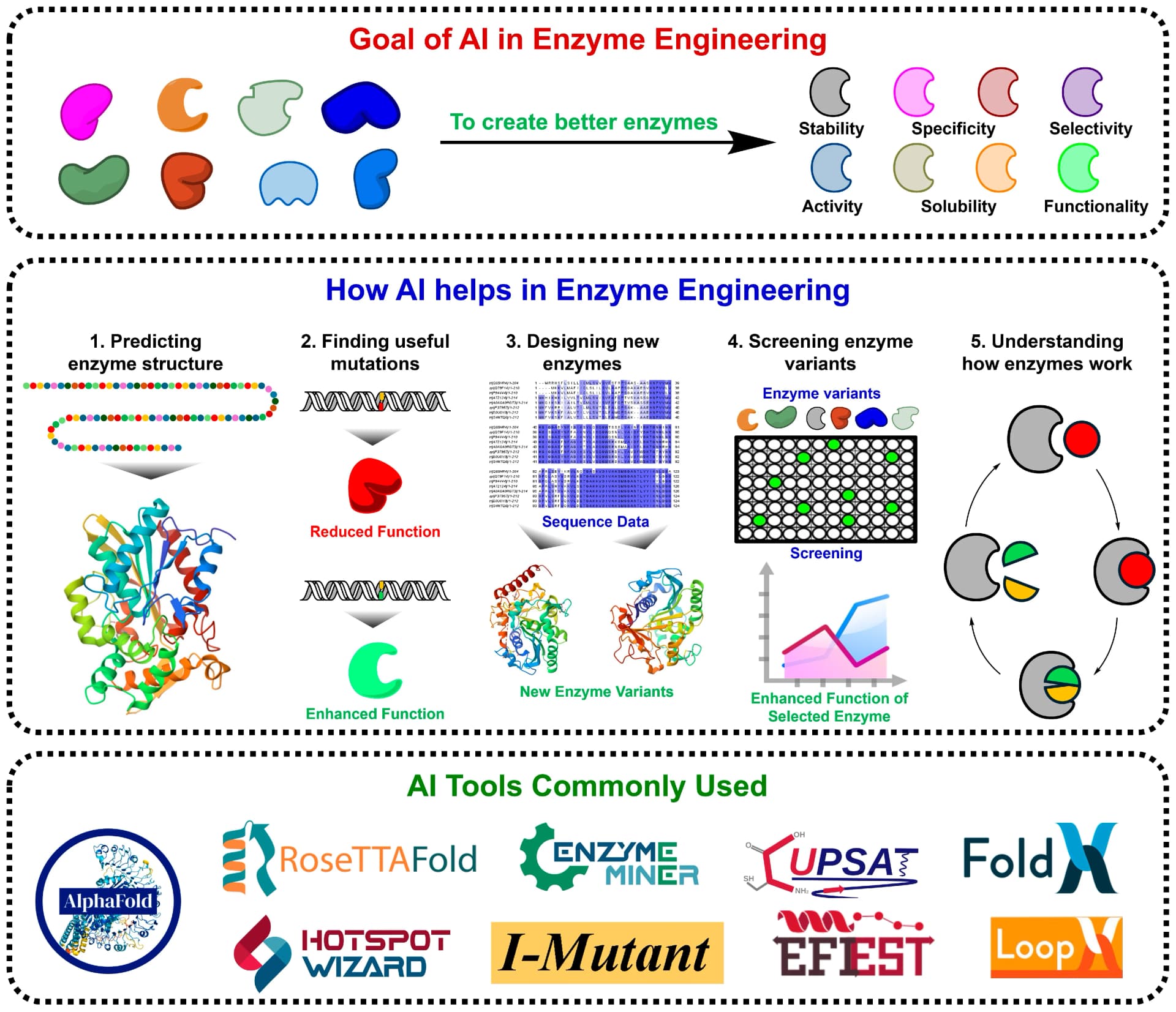
Researchers Mohd Faheem Khan (University College Dublin) and Mohd Tasleem Khan (Heriot-Watt University) set out to map the full landscape of AI methods now changing that picture. Their review in Molecules covers machine learning, deep learning, generative models, and reinforcement learning as applied to biocatalyst design, framing the field around a five-step workflow: structure prediction, mutation identification, variant design, functional screening, and mechanistic interpretation.
The scope spans pharmaceuticals, biofuels, detergents, food processing, and environmental remediation—anywhere enzymes matter industrially. One practical goal of the review was to give other researchers a curated toolkit they could act on, which is where ProteinIQ enters the story.
ProteinIQ in action
A key contribution of the review is Table 1, a comprehensive reference of AI tools organized by stage in the enzyme engineering pipeline. Compiling a resource like this requires the authors to evaluate not just scientific performance but also how researchers can realistically access and run each tool. Under the molecular docking category, the authors listed GNINA and PocketFlow as two tools researchers should have in their workflow, citing ProteinIQ as the web interface for both alongside their respective GitHub repositories.
Molecular docking is often the step that determines whether a proposed enzyme variant is worth building. Before committing to a mutagenesis campaign or cell-free expression run, researchers need to know how a substrate or inhibitor sits inside the active site, how tightly it binds, and whether an engineered mutation improves or disrupts that interaction.
GNINA approaches this problem with a convolutional neural network-based scoring function trained on large sets of protein-ligand complexes. Where classical docking programs like AutoDock Vina rely on additive physics-based force fields, GNINA learns representations of favorable binding geometries directly from data, producing more accurate pose rankings for flexible ligands and edge cases where physics-based scoring tends to disagree with experimental affinities. The review explicitly names GNINA alongside DiffDock as frameworks that "assist in predicting substrate binding and ligand interactions" across the enzyme engineering pipeline.
PocketFlow solves a problem that comes before docking: finding the binding pocket itself. For well-characterized enzymes, pocket location is obvious from prior structures. For newly designed enzymes, de novo sequences, or distant homologs with no close structural precedent, the active site geometry isn't known in advance. PocketFlow, developed by Tencent AI Lab, combines AI-based pocket prediction with flexible docking in a single workflow. Researchers can start from a predicted structure and get substrate binding results without manually defining a search box or relying on homology-based pocket annotation.
By listing ProteinIQ as the recommended access point for both tools, the authors positioned the platform as a practical on-ramp for computational enzyme engineers who want to run these analyses without local installation.
Key findings
The review synthesized findings across dozens of AI enzyme engineering studies and found consistent, large-scale improvements over conventional approaches. Autonomous design platforms integrating large language models with laboratory automation achieved up to 90-fold increases in substrate specificity; ML-guided cell-free expression systems evaluated over 10,000 reactions and identified variants with up to 42-fold enhanced catalytic efficiency; reinforcement learning identified multi-point beta-lactamase mutants with 2,000-fold gains in catalytic rate.
In detailed case studies, AlphaFold2-guided engineering of the GH11 xylanase PjxA raised its temperature optimum from 50°C to 70°C and boosted specific activity by approximately 4.7-fold, while ML-guided transaminase design for sitagliptin synthesis achieved 99.95% enantiopurity and cut manufacturing waste by 19%.
The authors also identified a recurring problem across generative AI applications: many sequences that score well computationally fail to fold or function when synthesized, which they call "computational artefacts." Closing that gap through tighter integration of experimental feedback, physics-aware modelling, and uncertainty quantification is where the authors see the most pressing opportunity for the field.