Simulation
gmx_MMPBSA
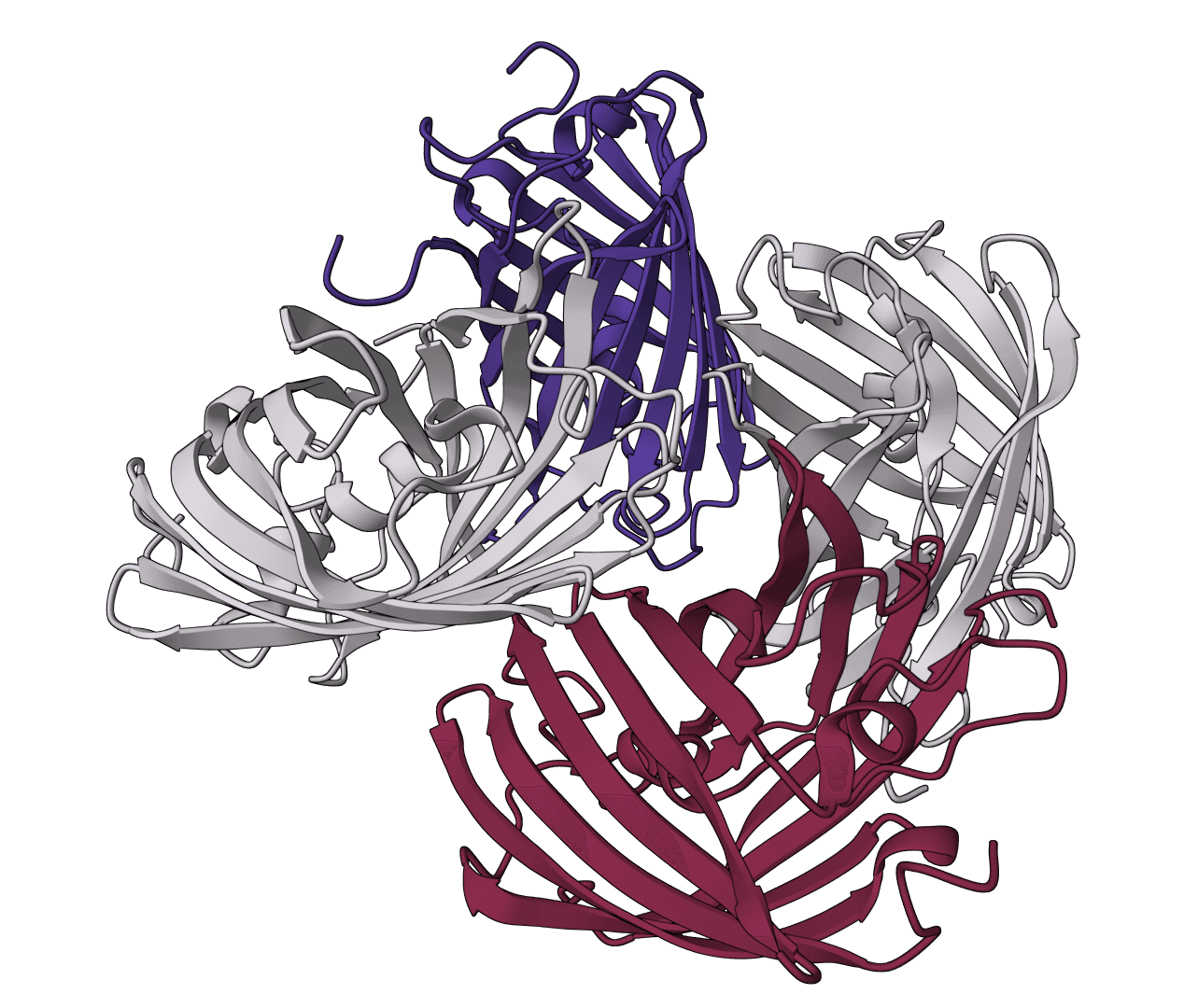
Calculate binding free energies using MM/PBSA and MM/GBSA methods for protein-ligand, protein-protein, and protein-DNA complexes. Provides detailed energy decomposition and per-residue contributions.

GROMACS
Run molecular dynamics simulations using the GROMACS engine with classical force fields (AMBER, CHARMM, GROMOS, OPLS). Study protein dynamics, conformational flexibility, and structural stability with production-grade MD methodology.
MD Trajectory Analysis
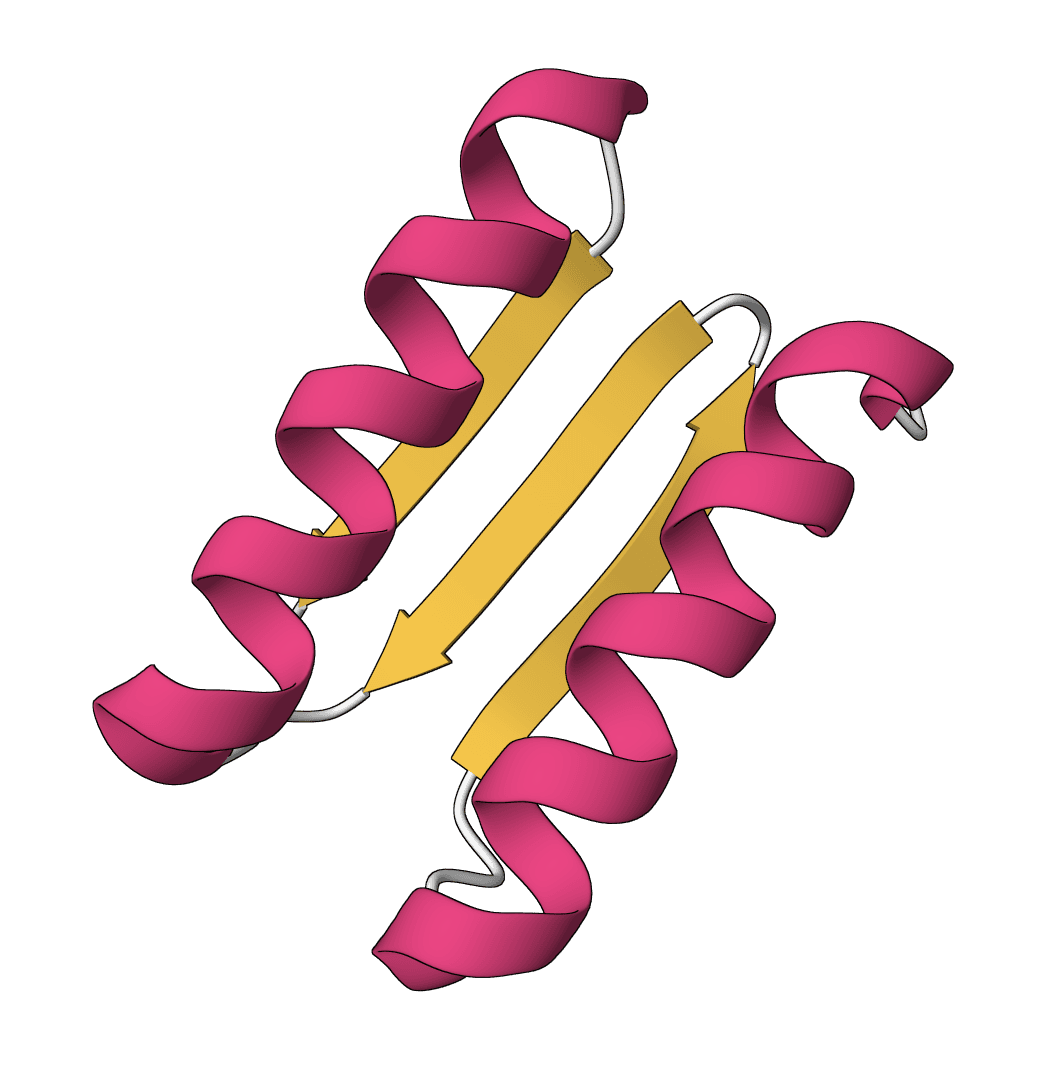
Analyze molecular dynamics trajectories using a ProteinIQ wrapper pinned to MDAnalysis 2.9.0. Calculate RMSD, residue-aggregated RMSF, radius of gyration, distance tracking, and additional trajectory observables from standard topology and trajectory files.
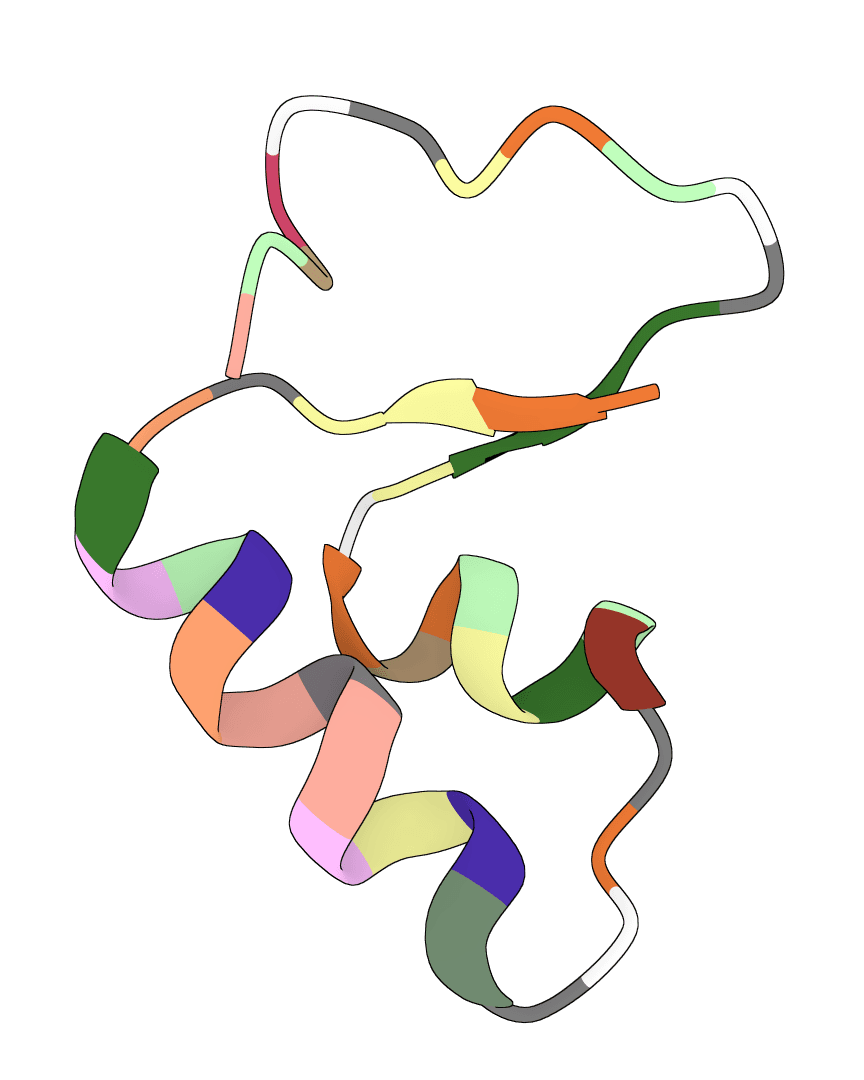
MDGen
MDGen is a generative AI model for molecular dynamics trajectory generation. Generate physically plausible conformational ensembles from a single protein structure, enabling rapid exploration of protein dynamics without expensive MD simulations.
OpenFE
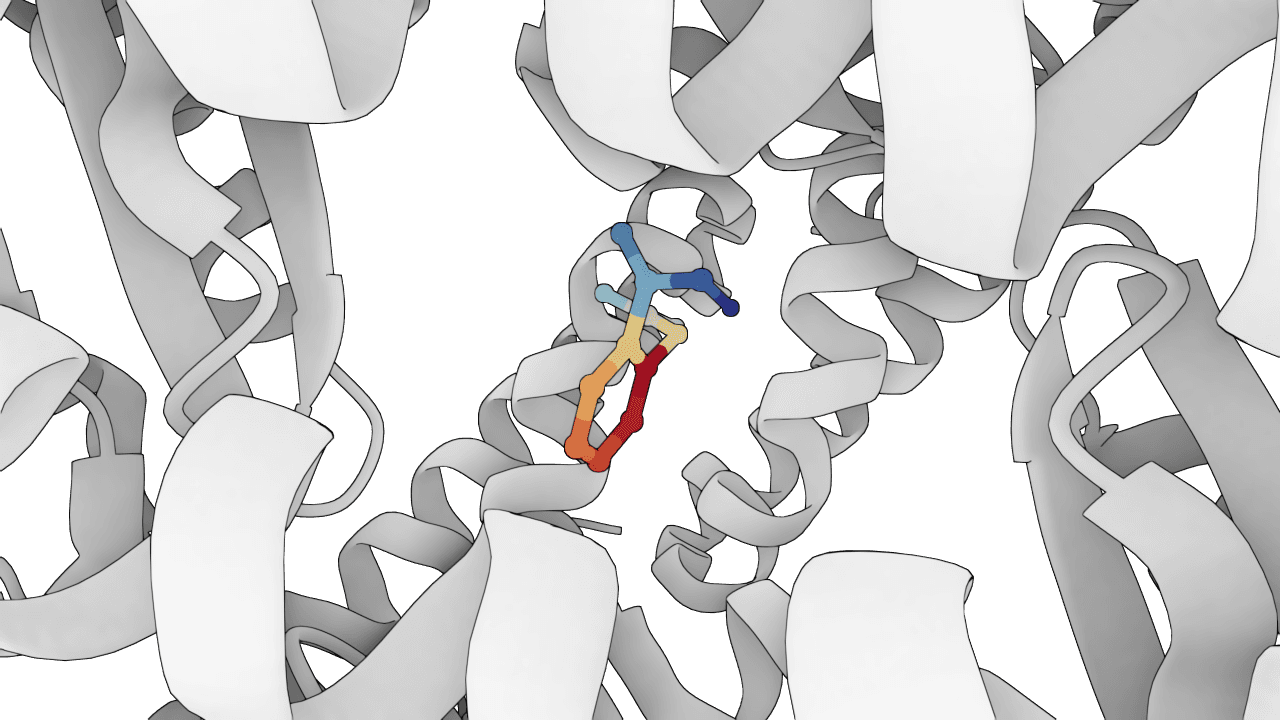
Run alchemical free energy calculations for drug discovery using Open Free Energy. Supports Absolute Hydration Free Energy (AHFE) and Relative Binding Free Energy (RBFE) calculations with GPU-accelerated OpenMM simulations.
OpenMM
Run GPU-accelerated molecular dynamics simulations using OpenMM. Simulate protein dynamics, study conformational changes, and analyze stability with industry-standard force fields (AMBER, CHARMM).
ORB v3
ORB v3 is a universal interatomic potential (machine learning force field) that predicts energies, forces, and stress tensors for atomic systems. Supports both molecular and materials structures with geometry optimization using conservative and direct model variants.