Screening
ADMET-AI
Predict ADMET (Absorption, Distribution, Metabolism, Excretion, Toxicity) properties from SMILES strings using machine learning models trained on Therapeutics Data Commons datasets.
Admetica
Predict 22 ADMET properties from SMILES strings with the upstream Admetica Chemprop models from Datagrok.

Brenk filter
Identify toxic, reactive, and pharmacokinetically problematic molecular fragments using structural alert patterns
DLKcat
DLKcat predicts enzyme turnover numbers (kcat values) from protein sequences and substrate structures using deep learning. Combines CNN and GNN architectures for accurate kinetic parameter prediction.
eToxPred
Predict toxicity and synthetic accessibility of small molecules using machine learning. eToxPred combines toxicity risk assessment with synthetic accessibility scoring to help prioritize drug candidates.
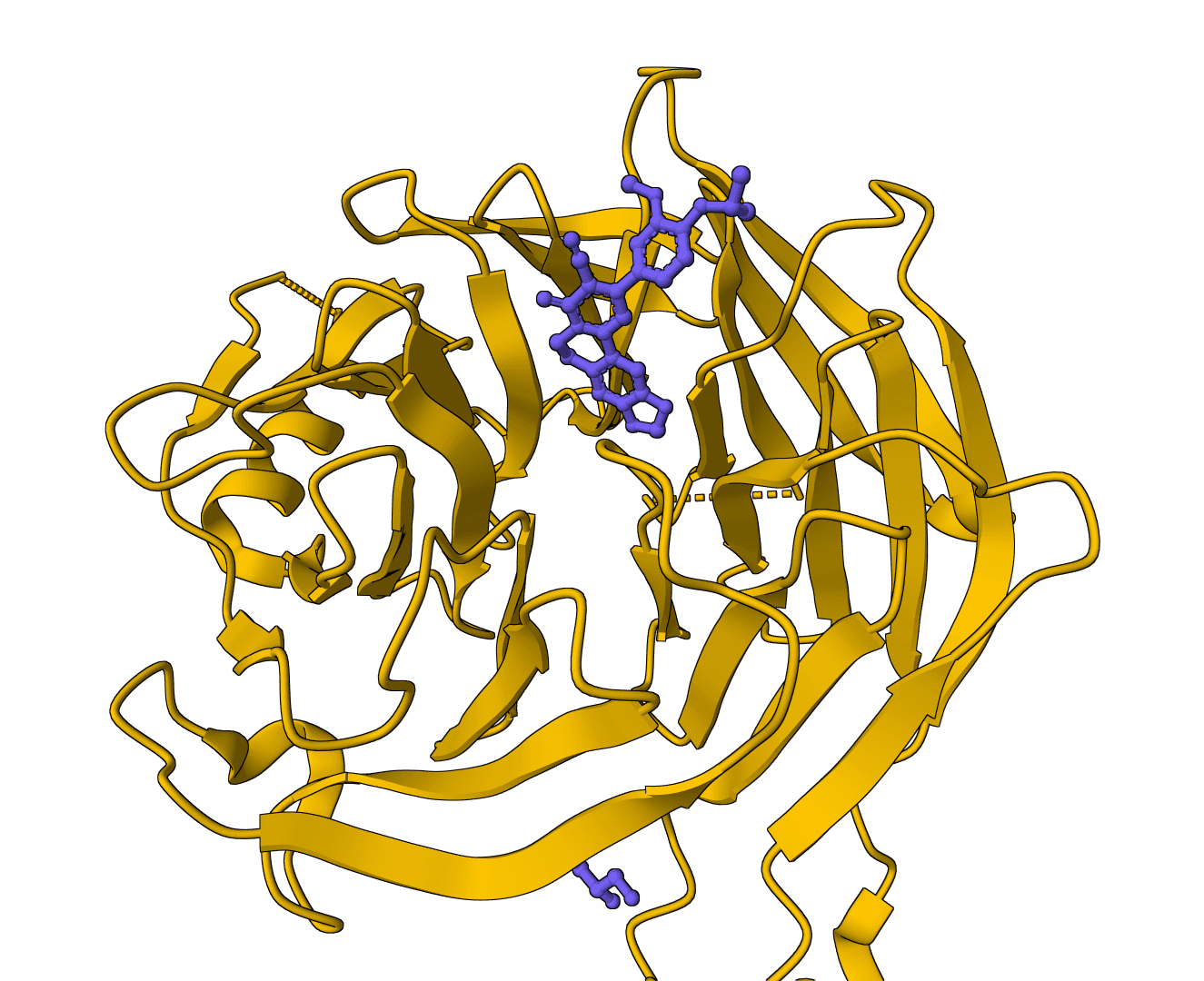
fpocket
Open-source protein pocket detection using Voronoi tessellation and alpha spheres. Identifies ligand binding sites with druggability scores.

Lead-likeness filter
Screen for lead-like compounds using stricter molecular descriptor criteria than Lipinski or Veber rules for early-stage drug discovery
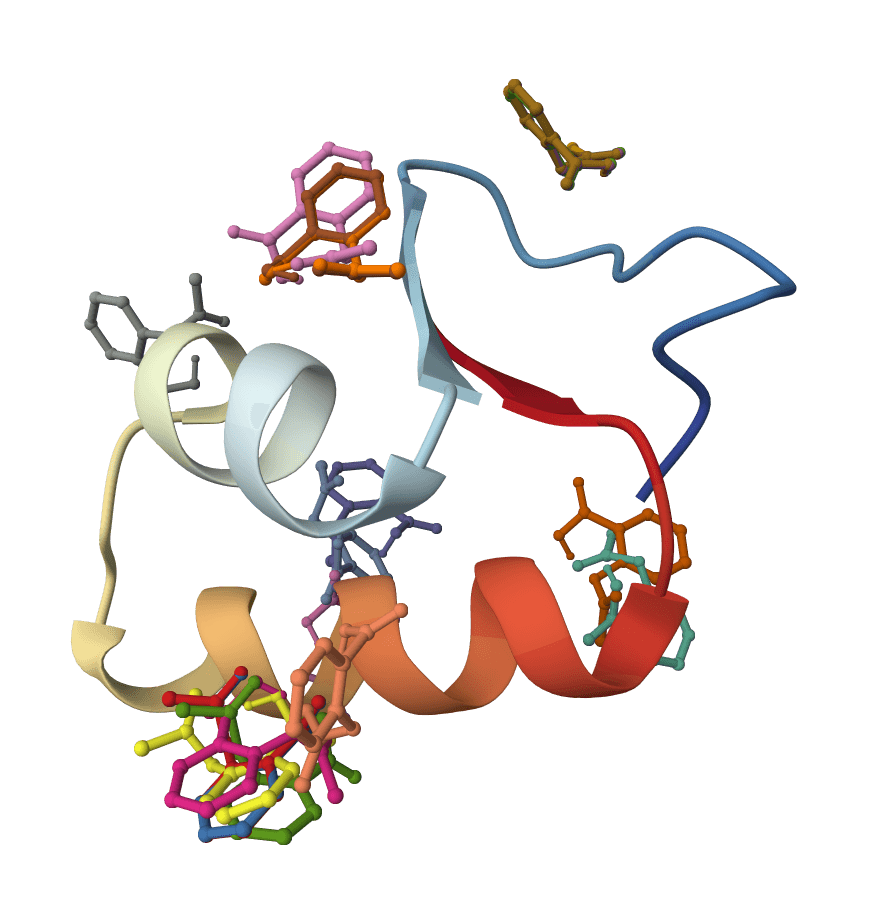
Lipinski's rule of 5
Lipinski's Rule of Five predicts whether compounds will be orally bioavailable by evaluating molecular weight, LogP, hydrogen bond donors, and acceptors.
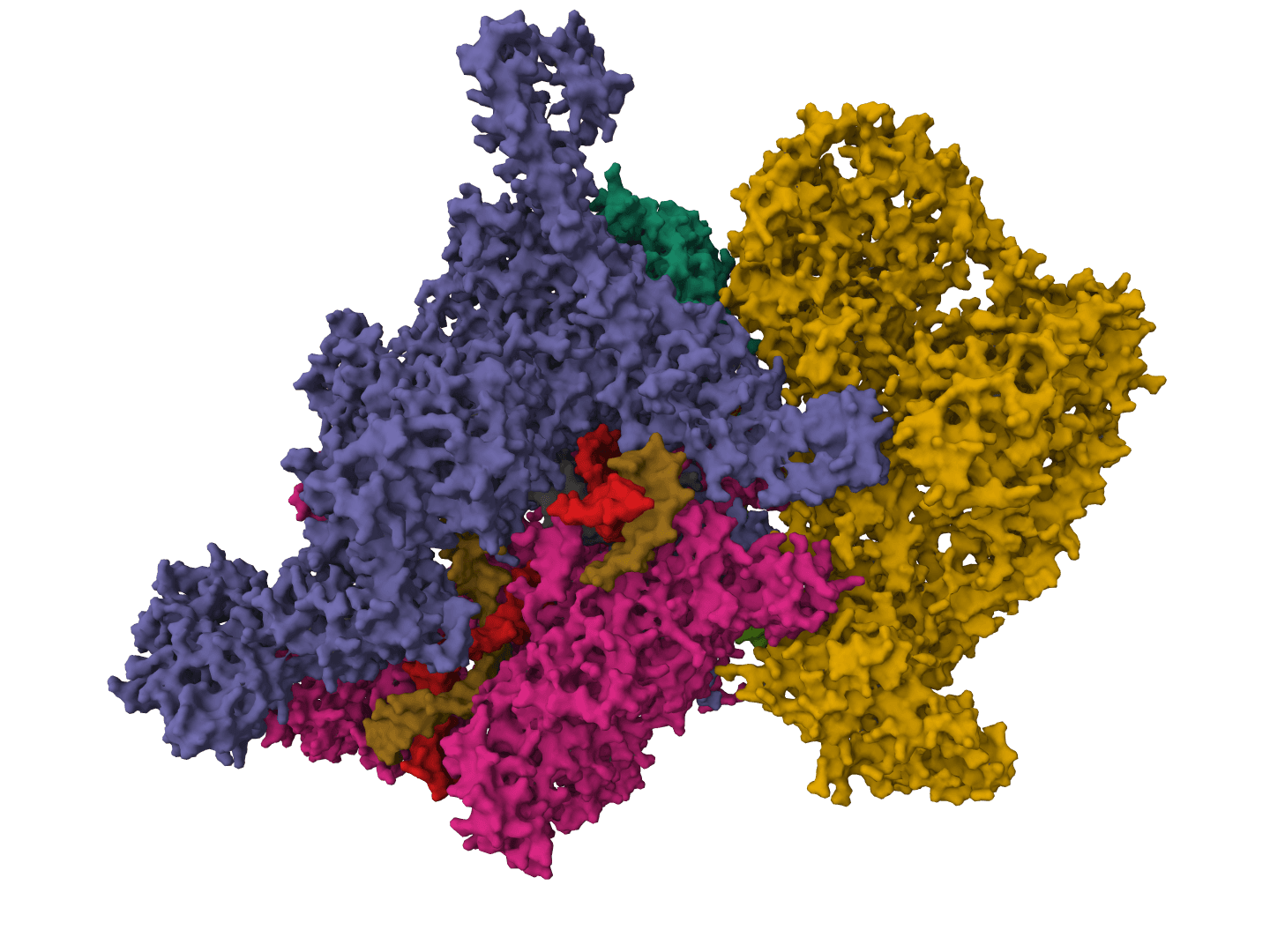
NetSolP-1.0
Predict protein solubility and usability for E. coli expression using ESM protein language models

PAINS filter
Screen compounds for Pan-Assay INterference patterns that cause false positives in biological assays

QEPPI
Quantitative estimate for protein-protein interaction inhibitor potential. Evaluates drug-likeness for compounds targeting PPIs.
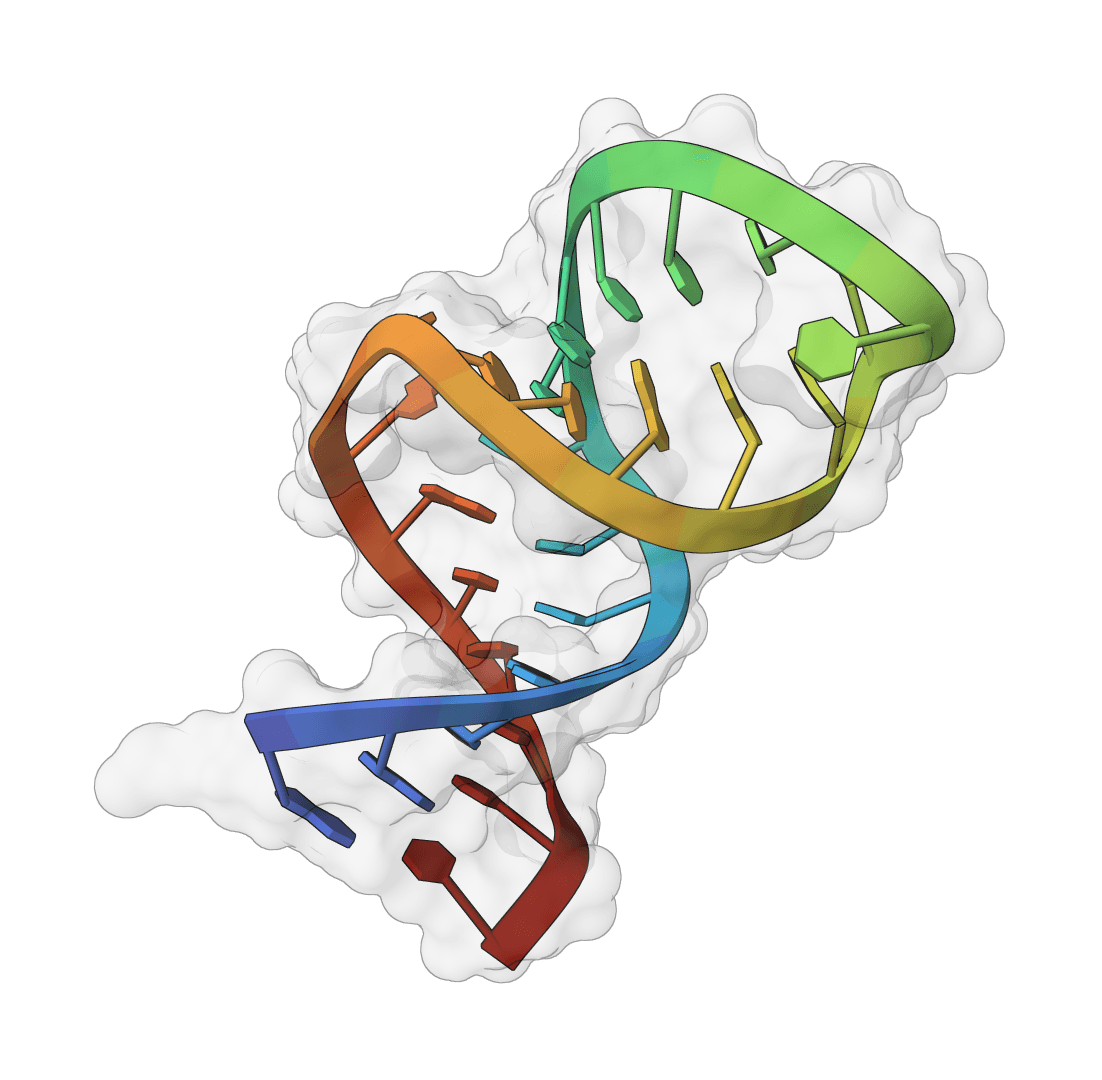
SMRTnet
Deep learning framework for predicting small molecule-RNA interactions using RNA secondary structure. Combines language models, CNNs, and graph attention networks for binding prediction.
SPRINT
SPRINT (Structure-aware PRotein ligand INTeraction) predicts drug-target interactions using co-embedded protein and ligand representations. Screen thousands of compounds against a protein target in seconds.
ToxPred 2.0 (Toxicity prediction)
Screen compounds for structural toxicity alerts using PAINS, Brenk, and NIH filters. For focused screening, see PAINS Filter, Brenk Filter, or Veber's Rule.

Veber's rule
Veber's Rule predicts oral bioavailability by evaluating molecular weight, LogP, hydrogen bond donors/acceptors, and rotatable bonds