Related tools
SPRINT
SPRINT (Structure-aware PRotein ligand INTeraction) predicts drug-target interactions using co-embedded protein and ligand representations. Screen thousands of compounds against a protein target in seconds.
ADMET-AI
Predict ADMET (Absorption, Distribution, Metabolism, Excretion, Toxicity) properties from SMILES strings using machine learning models trained on Therapeutics Data Commons datasets.
Admetica
Predict 22 ADMET properties from SMILES strings with the upstream Admetica Chemprop models from Datagrok.

Brenk filter
Identify toxic, reactive, and pharmacokinetically problematic molecular fragments using structural alert patterns
eToxPred
Predict toxicity and synthetic accessibility of small molecules using machine learning. eToxPred combines toxicity risk assessment with synthetic accessibility scoring to help prioritize drug candidates.

Lead-likeness filter
Screen for lead-like compounds using stricter molecular descriptor criteria than Lipinski or Veber rules for early-stage drug discovery
Lipinski's rule of 5
Lipinski's Rule of Five predicts whether compounds will be orally bioavailable by evaluating molecular weight, LogP, hydrogen bond donors, and acceptors.

PAINS filter
Screen compounds for Pan-Assay INterference patterns that cause false positives in biological assays

QEPPI
Quantitative estimate for protein-protein interaction inhibitor potential. Evaluates drug-likeness for compounds targeting PPIs.
ToxPred 2.0 (Toxicity prediction)
Screen compounds for structural toxicity alerts using PAINS, Brenk, and NIH filters. For focused screening, see PAINS Filter, Brenk Filter, or Veber's Rule.
What is SMRTnet?
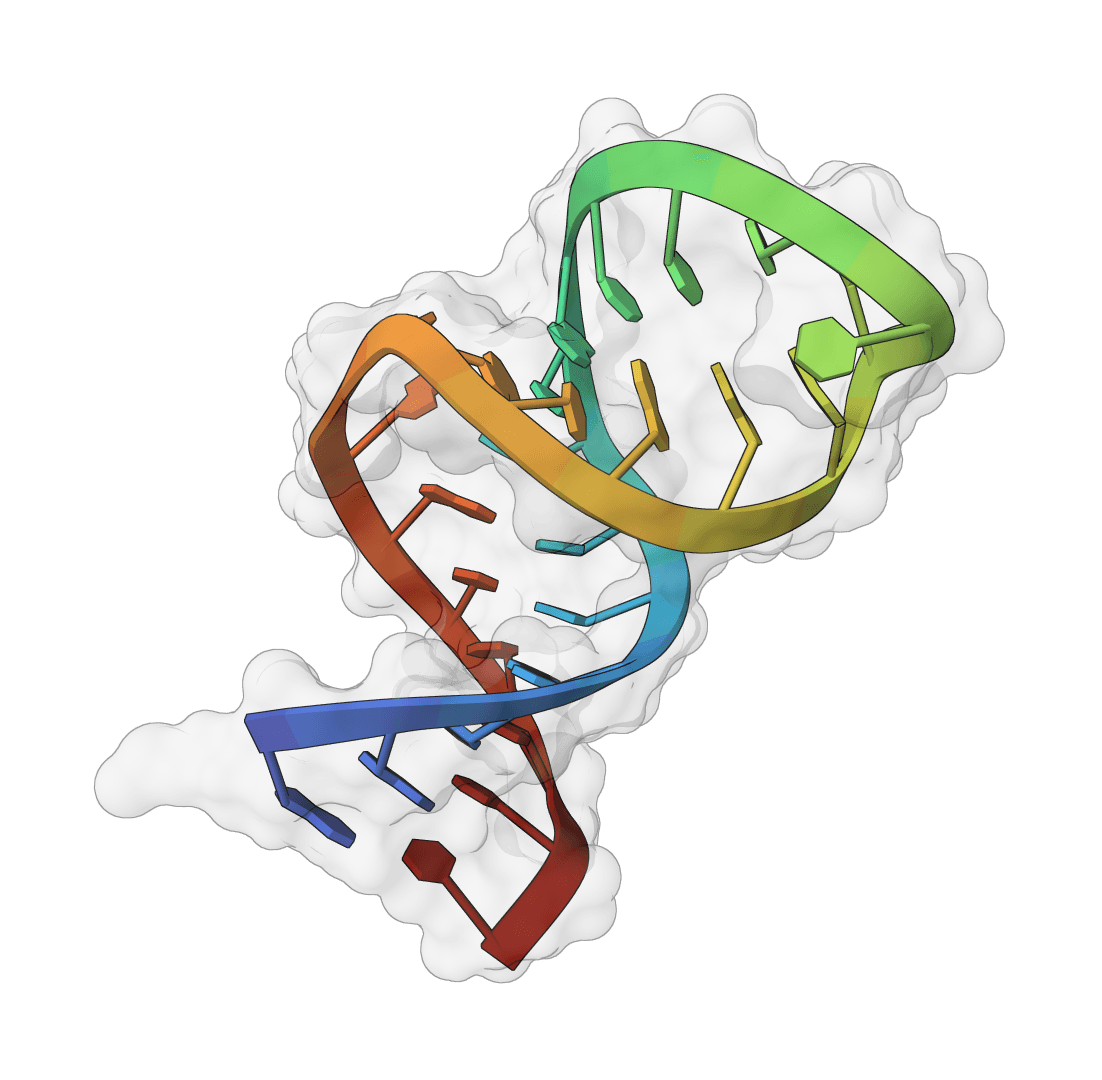
SMRTnet predicts whether a small molecule is likely to interact with an RNA target from RNA sequence, RNA secondary structure, and a molecular SMILES string.
The method is useful when the RNA target has no reliable 3D structure. Many disease-associated RNAs fall into that category: internal ribosome entry sites, untranslated regions, onco-miRNAs, viral RNA elements, and repeat expansion transcripts can have experimentally supported secondary structure while lacking a stable tertiary model suitable for docking.
SMRTnet was developed by Fei and colleagues for small molecule-RNA interaction prediction without RNA tertiary structures. The model combines RNA and chemical language models with CNN, graph attention, and multimodal fusion layers, then reports an ensemble binding probability for each RNA-compound pair.
How to use SMRTnet online
SMRTnet can be run on ProteinIQ by entering RNA targets as tab-separated name, sequence, and dot-bracket structure rows, then entering small molecules as tab-separated name and SMILES rows. The output is a ranked table of RNA-compound pairs with binding probability scores and binary interaction calls.
Inputs
| Input | Accepted format | Requirements |
|---|---|---|
RNA Target(s) | Text or .txt/.tsv file with three tab-separated columns: name, sequence, structure | RNA sequence and dot-bracket structure must have the same length. Minimum length is 31 nucleotides. Dot-bracket notation may use (, ), and .. |
Small Molecule(s) | Text or .txt/.tsv/.smi/.smiles file with two tab-separated columns: name, SMILES | SMILES must parse with RDKit. Invalid molecules are rejected rather than silently removed. |
RNA input should use RNA alphabet conventions, with U rather than T. Names are carried through to the output table, so short identifiers such as MYC_IRES, pre_miR_21, or SARS2_SL5 make results easier to review.
RNA input example
1MYC_IRES GUGGGGGCUUCGCCUCUGGCCCAGCCCUCAC (((((((((..(((...)))..)))))))))2TAU_UTR GCUAGCUAGCUAGCUAGCUAGCUAGCUAGCU ...(((((.......)))))............Small molecule input example
13902-71-4 CC1=CC(=O)OC2=C1C=C3C=C(OC3=C2C)C2caffeine CN1C=NC2=C1C(=O)N(C(=O)N2C)C3aspirin CC(=O)OC1=CC=CC=C1C(=O)OSettings
| Setting | Description |
|---|---|
Inference batch size | Number of RNA-compound pairs processed in each model inference batch. Default is 32; available values are 1, 4, 8, 16, 32, 64, 128, and 256. |
Inference batch size affects throughput and GPU memory use, not the scientific result. If a large screen fails because the batch does not fit in memory, lowering the value is appropriate. The same model weights, scoring logic, and interaction threshold are used.
Results
SMRTnet returns one row per RNA-compound pair, sorted by predicted binding probability.
| Column | Meaning |
|---|---|
Rank | Position after sorting by binding probability. 1 is the highest-scoring pair in the job. |
Molecule | Molecule name from the small molecule input. |
RNA Target | RNA target name from the RNA input. |
Binding Probability | Ensemble score from 0 to 1, rounded to four decimals. Higher values indicate stronger model support for interaction. |
Interaction | Yes when probability is at least 0.5; otherwise No. |
The output table is downloadable and copyable for follow-up triage. The ProteinIQ wrapper focuses on binding prediction and ranked screening results; it does not expose SMRTnet's separate attention-based binding site interpretation workflow.
Interpreting SMRTnet results
The binding probability is a model score, not a dissociation constant. A score of 0.82 means the ensemble strongly classifies the pair as an interaction relative to the training distribution. It does not mean 82 percent occupancy, 82 percent experimental success, or a specific value.
| Binding probability | Practical interpretation |
|---|---|
0.8 to 1.0 | Strong model support. These pairs are reasonable first candidates for MST, SPR, SHAPE-MaP perturbation, or reporter assays, especially when the RNA structure is experimentally supported. |
0.5 to 0.8 | Positive classification with moderate confidence. Useful for ranking analogs, expanding a hit list, or checking whether related molecules score consistently. |
0.3 to 0.5 | Below the default interaction threshold but not necessarily irrelevant. Consider these only when chemistry, target biology, or external evidence already supports the pair. |
0 to 0.3 | Low model support. These pairs are usually lower priority unless the input structure is uncertain or the molecule class is outside the model's training space. |
Score differences matter most near the top of a screen. For example, a set of natural products scoring 0.91, 0.88, and 0.86 against the same RNA should generally be treated as a top tier rather than as three precisely separated affinities. Conversely, a compound scoring 0.62 against one RNA and 0.18 against closely related decoy RNAs is more interesting than the same 0.62 score without selectivity context.
Secondary structure quality strongly affects interpretation. A structure supported by SHAPE-MaP, DMS probing, enzymatic probing, or a conserved motif model carries more weight than a single minimum-free-energy fold. For secondary structure prediction before screening, ViennaRNA can help generate dot-bracket hypotheses, but computational folds should be treated as hypotheses rather than confirmed binding-ready structures.
How SMRTnet works
SMRTnet is an upstream open-source deep learning method from the Zhang lab. ProteinIQ runs the published SMRTnet inference workflow rather than reimplementing the model.
The model uses four main representations:
- RNA sequence representation: RNASwan-seq, an RNA language model with transformer encoder blocks and rotary positional embeddings, learns sequence-level features from RNA tokens.
- RNA structure representation: A convolutional block reads dot-bracket secondary structure and extracts local pairing, loop, and stem-pattern features.
- Molecule language representation: MoLFormer converts the SMILES string into a learned chemical embedding.
- Molecule graph representation: A graph attention network represents the compound as atoms and bonds, then learns attention-weighted molecular features.
An attention-based multimodal fusion module combines RNA and molecule features before classification. For inference, SMRTnet uses five models from 5-fold cross-validation and aggregates their predictions with an ensemble strategy. ProteinIQ reports the resulting probability and classifies pairs at the 0.5 threshold used by the wrapper.
The original study evaluated SMRTnet on small molecule-RNA interaction benchmarks including R-BIND, R-SIM, SMMRNA, NALDB, and a NewPub set. The authors also tested predictions experimentally across 10 disease-associated RNA targets, including mRNAs of hard-to-drug proteins, onco-miRNAs, viral RNAs, and RNA repeat expansions. Forty hits were confirmed by microscale thermophoresis with nanomolar-to-micromolar dissociation constants, and one MYC IRES-associated compound showed downstream cellular effects in cancer cell lines.
When to use SMRTnet vs alternatives
SMRTnet is best for ligand-first or target-first screening against RNA when the available structural evidence is secondary structure. It is not a replacement for RNA-ligand docking when a high-quality 3D RNA structure and binding site are available.
| Method | Best fit | Main input requirement | Main caveat |
|---|---|---|---|
| SMRTnet | Ranking small molecules against RNA targets without tertiary structures | RNA sequence, dot-bracket secondary structure, SMILES | Returns interaction probability, not binding pose or affinity. |
| RNA-ligand docking | Pose generation and binding-site hypotheses for structured RNA pockets | 3D RNA structure and prepared ligand | Sensitive to RNA conformational flexibility and receptor preparation. |
| Experimental probing plus screening | High-confidence target validation and mechanism studies | Wet-lab structure or binding data | Slower and more expensive, but needed before biological claims. |
| Protein DTI models such as SPRINT | Protein target screening with small molecules | Protein sequence or structure-aware protein representation, SMILES | Designed for proteins, not RNA. |
A practical RNA-targeting workflow often starts with structure evidence, then screening, then validation. For example, a MYC IRES or viral stem-loop can be folded or constrained with experimental probing data, screened with SMRTnet, then followed by orthogonal assays on top-ranked molecules. If a candidate affects protein expression or viral replication, additional controls are needed to distinguish direct RNA binding from indirect cellular effects.
Limitations
- No binding pose: SMRTnet predicts interaction probability. It does not return a 3D RNA-ligand complex, docking pose, or atomic contact map in the ProteinIQ implementation.
- No affinity unit: Scores are not , IC50, or binding free energy. Experimental binding strength still needs measurement.
- Secondary structure dependence: Incorrect dot-bracket structures can move the model away from the biological RNA conformation of interest.
- Training distribution effects: Unusual chemotypes, modified nucleotides, protein-bound RNA conformations, and rare RNA motifs may score less reliably if similar examples were sparse in training data.
- Single-threshold classification:
Interactionuses a0.5probability cutoff. For real screening campaigns, ranking, replicate evidence, chemical tractability, and target selectivity usually matter more than a single yes/no label.