Related tools
PepMLM
Design linear peptide binders for target proteins using a target sequence-conditioned masked language model. PepMLM generates peptide sequences optimized to bind specific protein targets based on ESM-2 protein language modeling.
RFantibody
Structure-based de novo antibody and nanobody design pipeline combining antibody-tuned RFdiffusion, ProteinMPNN sequence design, and antibody-tuned RoseTTAFold2 filtering.
ODesign
All-atom generative AI for designing protein binders. Specify target binding sites and generate diverse binding proteins with fine-grained control over interaction parameters.
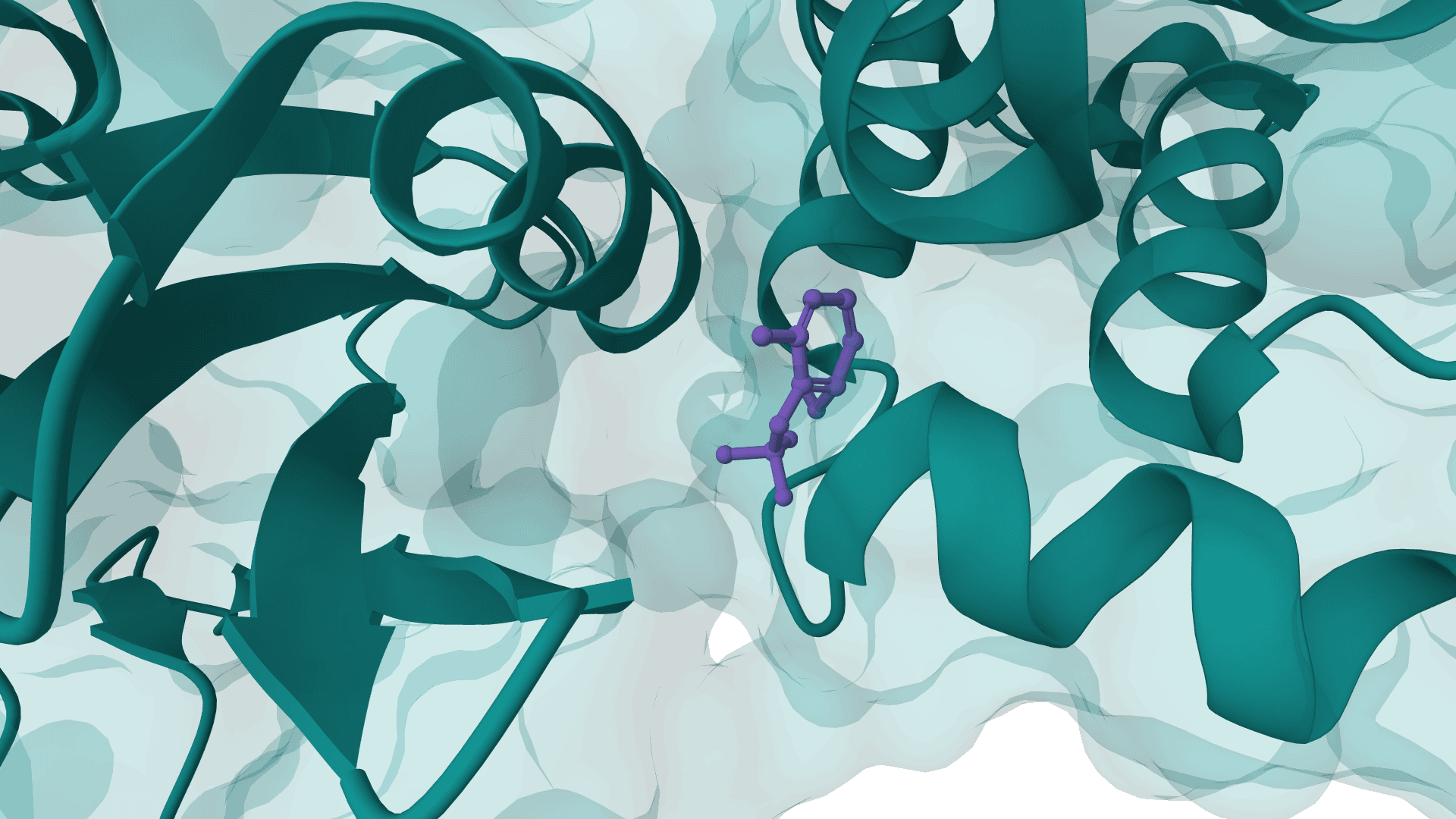
PocketFlow
PocketFlow is a structure-based molecular generative model that designs novel drug-like molecules within protein binding pockets. It uses autoregressive flow modeling with chemical knowledge to generate 100% chemically valid, highly drug-like compounds.
RFdiffusion
RFdiffusion is a state-of-the-art protein structure generation tool that uses diffusion models to design proteins de novo, create binders, scaffold motifs, and generate symmetric oligomers with atomic precision.
EvoDiff
EvoDiff is a diffusion-based protein sequence generation framework from Microsoft Research. ProteinIQ currently wraps the EvoDiff-Seq OA_DM_38M model for unconditional protein generation, motif scaffolding, and user-sequence inpainting.
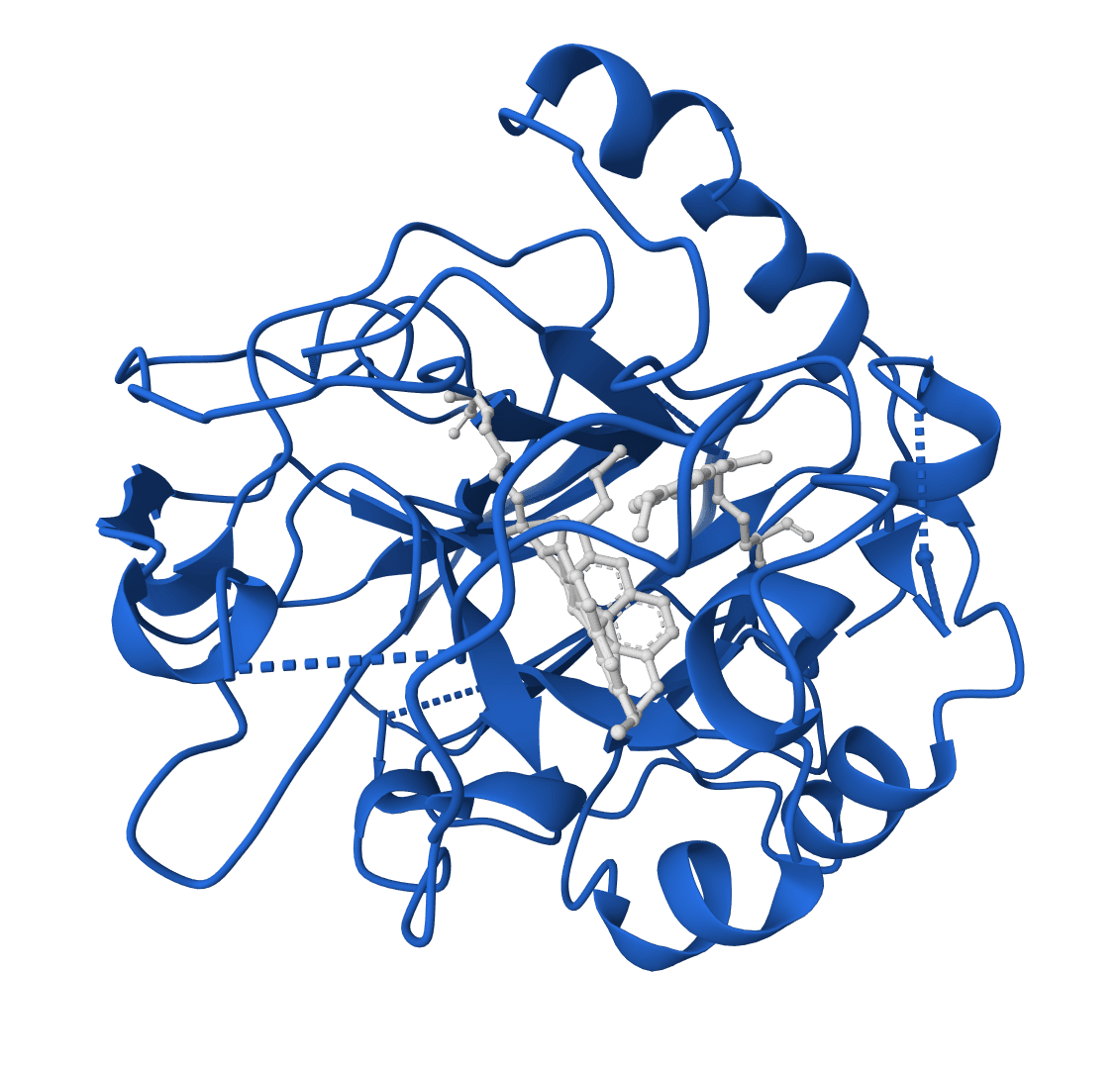
RFdiffusion 2
RFdiffusion2 is an atom-level enzyme active site scaffolding tool that generates protein scaffolds around your input motif. REQUIRES an input PDB structure containing the active site residues to scaffold. For ligand-aware design, ligands must be embedded in the input PDB as HETATM records.
BioPhi
Antibody humanization and humanness evaluation platform from Merck. Sapiens mode uses deep learning trained on the Observed Antibody Space (OAS) to humanize antibody sequences, while OASis mode evaluates humanness using 9-mer peptide search against human antibody databases.
GenMol
GenMol is a generative AI model from NVIDIA that creates novel drug-like molecules using masked discrete diffusion. It generates molecules in SAFE representation format and supports de novo generation, linker design, motif extension, and scaffold decoration.

ProFam
ProFam-1 is a protein family language model for family-conditioned sequence generation. Provide a protein family FASTA/MSA and generate new sequences with model likelihood scores for downstream ranking and screening.
What is BoltzGen?
BoltzGen is an all-atom diffusion model for binder design. It generates new peptides, miniproteins, nanobodies, and antibody/Fab CDR binders directly against a supplied target structure, and it also includes a protein-small molecule mode for designing proteins around a ligand.
Rather than optimizing a fixed starting sequence, BoltzGen samples binder sequence and structure together. That makes it useful when no obvious scaffold exists or when several geometrically distinct binding solutions are worth exploring. The model was introduced by Hannes Stärk and was experimentally evaluated across multiple wet-lab campaigns, with especially strong results on novel targets where template-heavy approaches tend to struggle.
How to use BoltzGen online
ProteinIQ runs BoltzGen on hosted GPU infrastructure, so binder design jobs can be configured in the browser and submitted without setting up CUDA, PyTorch, or the BoltzGen CLI. The online interface exposes the current v0.3.0 protocol family and the ranking controls that were added in newer BoltzGen releases.
Inputs
| Input | Description |
|---|---|
Target protein | Upload a .pdb, .ent, .cif, or .mmcif structure file, or fetch a structure from the RCSB PDB by ID. Used for Peptide, Protein, Nanobody, and Antibody / Fab CDR protocols. |
Target ligand | Used for Protein-small molecule mode. Accepts a SMILES string, CCD code, or PubChem lookup. |
Job name | Optional label for tracking runs in ProteinIQ job history. |
Settings
Core settings
| Setting | Description |
|---|---|
Protocol | Design mode. Peptide is intended for short binders, Protein for de novo miniproteins, Nanobody for single-domain antibody binders, Antibody / Fab CDR for scaffold-guided antibody CDR design, and Protein-small molecule for designing a protein around a ligand. |
Antibody scaffold | Available for Antibody / Fab CDR. ProteinIQ includes therapeutic antibody scaffolds such as adalimumab, dupilumab, and ustekinumab, or All scaffolds for broader sampling. |
Number of designs | Total number of candidates generated before filtering. Runtime and credit cost scale with this value. |
Budget | Number of final designs retained after BoltzGen applies quality and diversity filtering. |
Binder settings
| Setting | Description |
|---|---|
Uniform binder size | Uses a single exact binder length when enabled. Disabling it allows length sampling between the configured minimum and maximum. |
Binder length | Target binder length when uniform sizing is enabled. Appropriate ranges depend on protocol: peptides are short, protein binders are larger, and nanobody / antibody modes use scaffold-driven sizes. |
Minimum length | Lower bound for sampled binder size when Uniform binder size is off. |
Maximum length | Upper bound for sampled binder size when Uniform binder size is off. |
Binding site | Optional target-site constraint in chain:residues form, for example A:12,14,61. This biases designs toward a specified epitope or pocket. |
Cyclic peptide | Adds an N-to-C terminal cyclization constraint for peptide binders. |
Target chains | Optional comma-separated subset of target chains to keep, for example A,B. |
Inverse folding
| Setting | Description |
|---|---|
Skip inverse folding | Stops after backbone generation and skips sequence redesign. |
Sequences per backbone | Number of inverse-folded sequences produced for each designed backbone. |
Avoid amino acids | One-letter amino acid codes to exclude during inverse folding, such as C to avoid undesired disulfides. |
Filtering and ranking
| Setting | Description |
|---|---|
Quality vs diversity (alpha) | Controls the tradeoff between top-scoring designs and structural diversity in the final ranked set. |
Filter biased compositions | Removes amino-acid composition outliers. The current upstream default is true, and ProteinIQ matches that behavior. |
Refolding RMSD threshold | Upper RMSD cutoff used during refolding-based filtering. Most relevant for protein-sized binders. |
Custom filters | Extra hard filters in metric>value or metric<value form, one per line. |
Metrics weights | Per-metric ranking weights using the current metric=value syntax, one per line. Larger values down-weight a metric rank, and metric=none removes that metric from ranking. |
Size buckets | Optional cap on how many retained designs may come from each size range. Format: min-max:count, one per line, for example 10-20:5. This is useful when sampling variable binder lengths. |
Diffusion sampling
| Setting | Description |
|---|---|
Step scale | Diffusion step size. Higher values generally increase exploration at the cost of stability. |
Noise scale | Noise level during sampling. Lower values make generations more deterministic. |
Model checkpoint | Both mixes BoltzGen's diverse and adherence checkpoints. Diverse favors novelty, while Adherence favors constraint fidelity. |
Structure constraints
| Setting | Description |
|---|---|
Secondary structure | Binder secondary-structure constraints in chain:start-end:type format, one per line. Supported types are HELIX, SHEET, and LOOP. |
Disulfide bonds | Cysteine bridge constraints in chain:residue,chain:residue format. |
Staple bonds | Non-natural crosslinks in chain:residue:atom,chain:residue:atom format. |
Advanced design
| Setting | Description |
|---|---|
Fixed sequence regions | Locks a binder segment to a specific sequence using chain:start-end:sequence. |
Binding residues | Declares binder positions that should contact the target. |
Non-binding residues | Declares binder positions that should avoid target contact. |
Design insertions | Variable-length insertion syntax in chain:position:min..max form. The field is exposed in the interface, but the current ProteinIQ BoltzGen integration does not yet apply these insertions during YAML generation. |
Execution control
| Setting | Description |
|---|---|
Reuse existing results | Reuses compatible intermediate outputs if the same run directory is resumed. |
Pipeline steps | Runs a specific stage such as design, inverse_folding, folding, analysis, or filtering instead of the full pipeline. |
Redesign existing structure | Equivalent to BoltzGen's inverse-fold-only mode. Requires a fully specified structure with the binder already positioned. |
Results
ProteinIQ returns ranked designs in an interactive viewer together with downloadable structure and sequence files. Runs also include the original uploaded reference input so the designed binders can be compared against the starting target or ligand.
| Output | Description |
|---|---|
Viewer | Interactive structural view of ranked designs and reference inputs. |
Rank | Position in the final filtered list. Lower ranks are preferred. |
Quality (pTM) | Predicted complex quality score on a 0 to 1 scale. Higher values indicate stronger structural confidence. |
Error (Å) | Predicted aligned error for the complex. Lower values indicate a more confident model. |
Interface (Ų) | Buried surface area between binder and target. Larger interfaces often indicate more extensive contacts, though optimal values depend on binder class. |
Sequence | Designed amino acid sequence for the retained candidate. |
Files | Downloadable CIF structures, FASTA sequences, and reference inputs when available. |
Interpreting results
Quality (pTM)
| Range | Interpretation |
|---|---|
0.8-1.0 | Strong structural confidence. Often the first tier to inspect experimentally. |
0.6-0.79 | Usable designs with moderate confidence. Visual inspection and orthogonal validation are advisable. |
<0.6 | Lower-confidence models that may still be interesting for difficult targets, but usually require more screening. |
Error (Å)
| Range | Interpretation |
|---|---|
<3 Å | High-confidence binder-target geometry. |
3-5 Å | Intermediate confidence. Interfaces may still be plausible but should be reviewed. |
>5 Å | Lower-confidence complexes or flexible interfaces. |
How does BoltzGen work?
All-atom diffusion
BoltzGen models the binder and target at all-atom resolution instead of working only with a backbone trace. The design process begins from noisy coordinates and repeatedly denoises them while conditioning on the target geometry and any user-supplied constraints. Sequence identity is coupled to the structural representation, so side-chain arrangement and residue type are learned together rather than stitched together as separate steps.
Protocol families
The same framework supports several design regimes:
- Peptide binders: Short linear or cyclic peptides for compact interfaces and pockets
- Protein binders: De novo miniproteins for larger or more structured interaction surfaces
- Nanobody binders: Single-domain antibody designs with nanobody-style geometry
- Antibody / Fab CDR design: CDR generation on fixed therapeutic antibody scaffolds
- Protein-small molecule design: Protein design around a ligand, with additional affinity-oriented analysis
Pipeline execution
In ProteinIQ, BoltzGen typically runs the same five major stages exposed by the upstream CLI:
design: Generates candidate backbonesinverse_folding: Assigns or redesigns sequencesfolding: Refolds designed candidates for structural validationanalysis: Computes interface and quality metricsfiltering: Selects the final set by balancing quality and diversity
Protein-small molecule runs can add ligand-specific analysis, and Redesign existing structure skips backbone generation and enters directly at inverse folding.
Constraint language
BoltzGen's design specification language makes it possible to bias generation without retraining the model:
- Binding-site constraints: Focus contact formation on specific target residues
- Secondary-structure constraints: Favor helices, sheets, or loops in selected binder segments
- Covalent constraints: Encode cyclic peptides, disulfides, and staples
- Sequence constraints: Keep selected residues fixed or bias interaction positions
These constraints do not guarantee success, but they substantially narrow the search space when a design hypothesis already exists.
Limitations
- Target flexibility is limited: Large induced-fit rearrangements are not modeled explicitly during design.
- Cost grows quickly with binder size: Protein-length binders are substantially more expensive than short peptides.
- Experimental success is still context-dependent: High pTM and low PAE do not guarantee expression, solubility, or binding in vitro.
- Protein-small molecule design is more specialized: It is useful for ligand-focused problems, but the main BoltzGen validation literature emphasizes peptide, nanobody, and antibody-style binder design.
Design insertionsare not active in the current ProteinIQ integration: The field is visible in the UI, but those insertions are not yet emitted into the generated BoltzGen YAML.- Inverse-fold-only mode requires a prepared structure: The binder must already be positioned in the uploaded complex if
Redesign existing structureis enabled.