What is Chai-1?
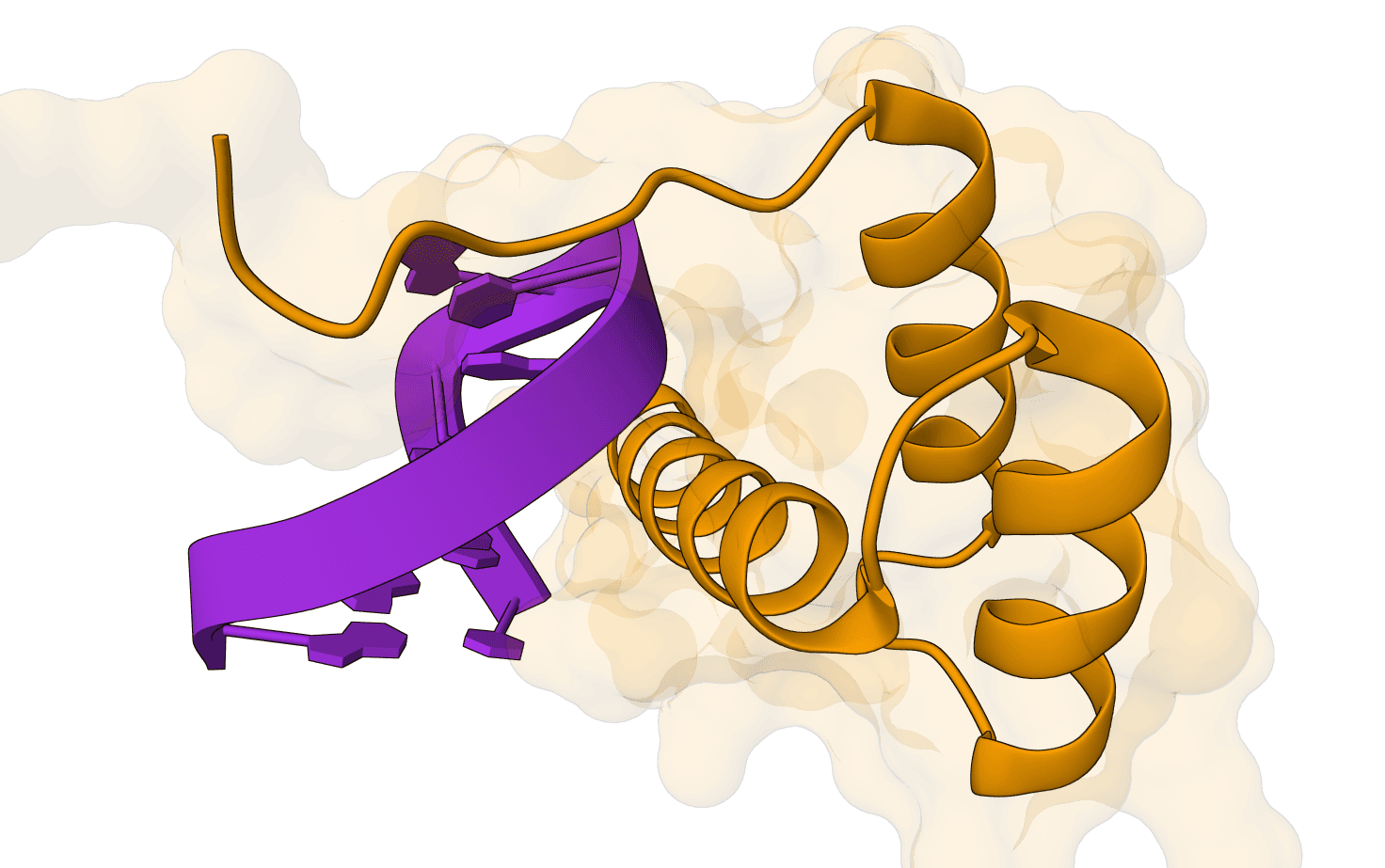
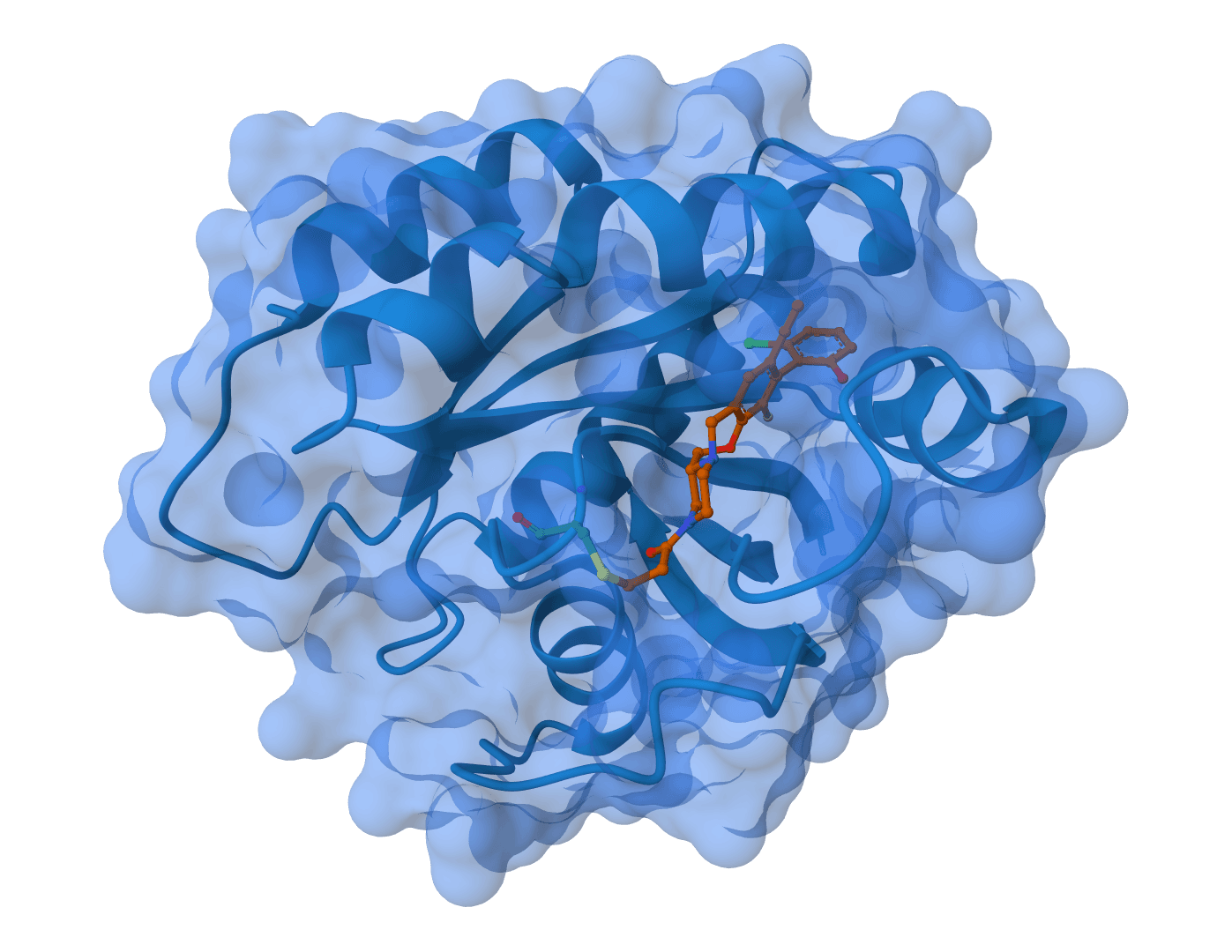
Chai-1 is a multi-modal foundation model for molecular structure prediction developed by Chai Discovery. Released in October 2024, it can predict 3D structures for proteins, small molecules, DNA, RNA, and multi-component complexes containing any combination of these molecular types.
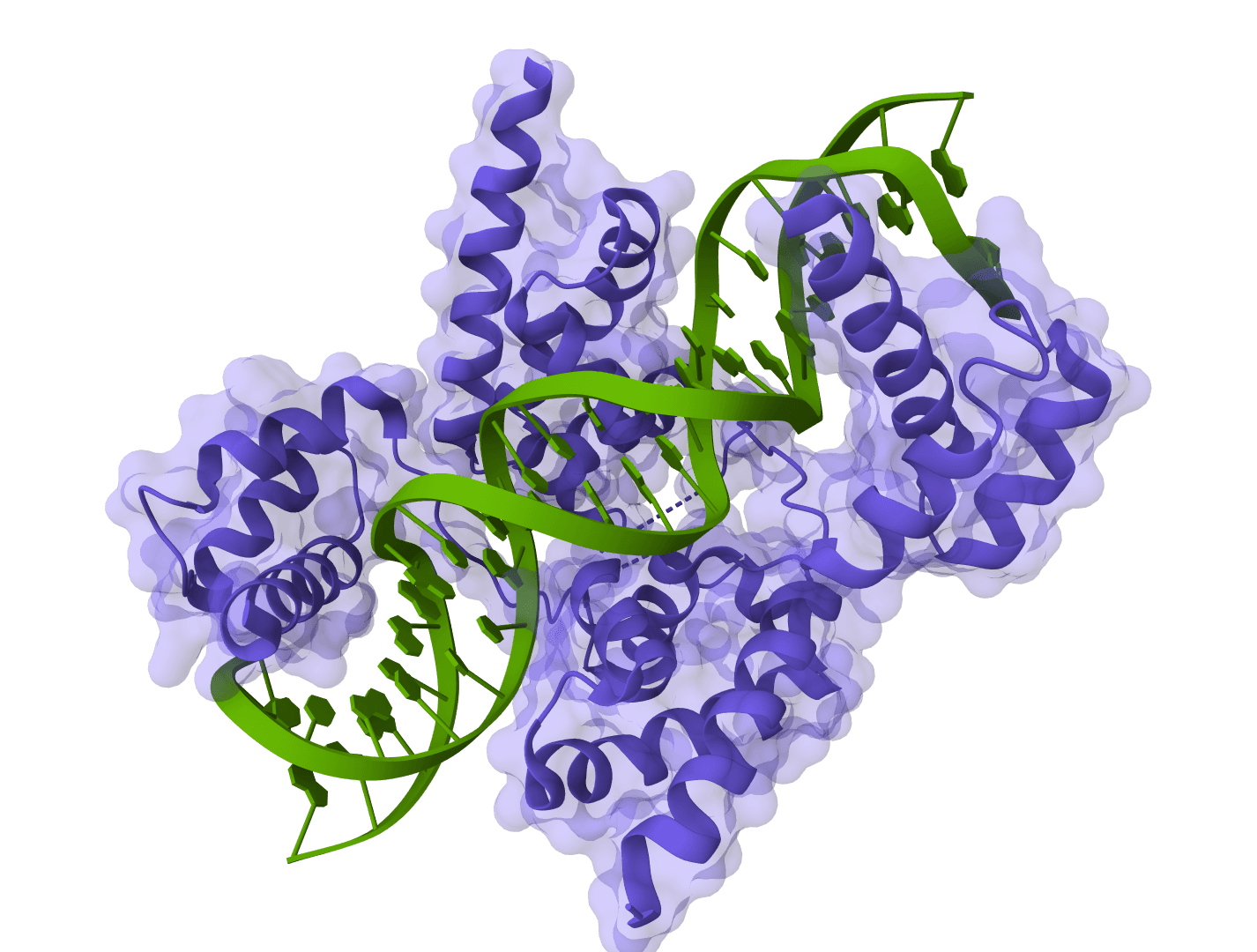
Chai-1 addresses a key limitation of earlier structure prediction tools by supporting truly unified multi-modal predictions. Rather than requiring separate models for different molecular types, Chai-1 handles everything in a single framework—including protein-ligand complexes, protein-DNA interactions, and systems with post-translational modifications.
Since its relase, the Chai-1 performance and capabilities have been reached and exceded by Boltz-2 and BoltzGen.
How does Chai-1 work?
Chai-1 uses a diffusion-based architecture to generate molecular structures. Like image generation models, it starts from random noise and iteratively refines toward physically plausible 3D coordinates through learned denoising steps.
Diffusion-based structure generation
The model learns to reverse a diffusion process that gradually corrupts molecular structures with noise. During inference, it starts from random atomic coordinates and progressively denoises them through hundreds of steps to produce a coherent structure.
Each diffusion step adjusts atomic positions based on the model's learned understanding of molecular geometry, chemical bonding, and inter-molecular interactions. More diffusion steps generally produce higher-quality structures at the cost of increased computation time.
Evolutionary information from MSA
For protein structure prediction, Multiple Sequence Alignment (MSA) provides crucial evolutionary information. MSA aligns your target protein against thousands of homologous sequences from different organisms.
Coevolution patterns in the MSA reveal which residues are spatially close in 3D—if two positions consistently mutate together across evolution, they're likely in contact. This signal is one of the most powerful inputs for modern structure prediction.
Chai-1 can automatically generate MSAs via the ColabFold server, searching sequence databases to find homologs for each protein chain.
Recycling for iterative refinement
Chai-1 uses a recycling mechanism where the model feeds its prediction back through the network multiple times. Each recycling step allows the model to refine its prediction based on the previous iteration, progressively improving accuracy.
This is similar to how AlphaFold2 works—the model essentially "thinks twice" about difficult regions, using information from earlier predictions to resolve ambiguities.
Multi-modal representation
Unlike models trained exclusively on proteins, Chai-1 was designed from the ground up to handle diverse molecular types. Proteins, nucleic acids, and small molecules are all represented in a unified framework that captures their distinct chemical properties while enabling cross-modal interactions.
This unified architecture is what enables Chai-1 to predict complex multi-component systems like protein-ligand complexes, protein-DNA interactions, or systems with covalently attached modifications.
Molecule inputs
Chai-1 accepts multiple molecule types that you can combine into a single prediction job. Chain IDs (A, B, C...) are assigned automatically and displayed in the UI—you'll need these when defining constraints.
- Protein: Upload PDB/CIF files, paste FASTA sequences, or fetch directly from RCSB using a PDB ID. Up to 10 chains.
- Ligand (SMILES): Enter SMILES strings, upload SDF/MOL/MOL2 files, or fetch from PubChem by compound ID. Up to 10 ligands.
- DNA: Paste sequences in FASTA format or upload structure files. Up to 10 chains.
- RNA: Paste sequences in FASTA format or upload structure files. Up to 10 chains.
Prediction parameters
- Number of samples: How many structure predictions to generate (1–10). More samples provide conformational diversity but increase runtime.
- Recycling steps: Number of refinement iterations where the model feeds its prediction back through the network. More steps improve accuracy but increase runtime.
- Diffusion steps: Number of denoising steps in the diffusion process. Higher values produce more refined structures at the cost of speed. We recommend 200 for routine predictions and 300–500 for high-accuracy needs.
- Random seed: Fixed seed for reproducible predictions. Leave empty for random seed each run.
MSA & templates
MSA (Multiple Sequence Alignment) provides evolutionary context that substantially improves protein structure prediction accuracy. Templates are known experimental structures from homologous proteins that can guide the prediction.
- Generate MSA: Enables automatic MSA generation via the ColabFold server. When enabled, Chai-1 searches sequence databases to find homologous sequences for each protein chain. Disable this only for quick screening runs or when working with synthetic/designed proteins that lack natural homologs.
- Use templates: Enables structural template search from the PDB. Templates provide experimental structural context from homologous proteins. We recommend enabling this for difficult targets or when high accuracy is critical.
Constraints
Constraints guide the prediction toward specific structural features when you have prior knowledge about the system. This is particularly useful when you know the binding site from experimental data or have crosslinking mass spectrometry data.
- Contact constraints: Forces two residues to remain within a specified distance. Format:
chain:residue,chain:residue|max_distance. Example: A:10,B:5|8.0 constrains residue 10 of chain A within 8Å of residue 5 in chain B.
- Pocket constraints: Forces a ligand to bind near specified residues. Use this when the binding site is known from crystallography or mutagenesis. Format:
ligand_chain|protein:res1,protein:res2|max_distance. Example: C|A:45,A:46|6.0 constrains chain C (ligand) near residues 45–46 of chain A (protein).
Modifications
Post-translational modifications (PTMs) can significantly affect protein structure. Chai-1 supports common modifications using standard CCD (Chemical Component Dictionary) codes from the PDB.
- Modified residues: Specifies modified residues using chain IDs shown in the molecule labels. Format:
chain:position:CCD_code. Common codes: SEP (phosphoserine), TPO (phosphothreonine), PTR (phosphotyrosine), MSE (selenomethionine), MLY (methylated lysine).
Understanding the results
Chai-1 returns multiple ranked predictions, each with a structure file and associated confidence information.
Structure files
Each prediction produces a CIF structure file representing one possible conformation. We recommend examining the top 2–3 structures rather than relying solely on the highest-ranked prediction—especially for flexible complexes or multi-domain proteins.
Confidence interpretation
If predictions differ substantially across samples, this may indicate:
- The target is intrinsically flexible
- The model is uncertain about the conformation
- MSA depth is insufficient for confident prediction
In such cases, consider enabling MSA generation or increasing the number of samples to better explore conformational space.
Best practices
Getting good predictions
Enable MSA for proteins. The evolutionary information from MSA substantially improves prediction accuracy. Only disable it when working with synthetic proteins that lack natural homologs, or when you need a quick rough estimate.
Use experimental structures when available. If you have an experimental structure for part of your complex (e.g., the protein receptor), provide it directly rather than predicting from sequence.
Check your SMILES stereochemistry. Ligand binding is highly stereospecific. Make sure your SMILES strings correctly represent the enantiomer you intend to model.
Choosing the right settings
| Use case | Samples | MSA | Diffusion steps | Notes |
|---|
| Quick screening | 1–3 | Off | 100–150 | Fast turnaround, lower accuracy |
| Standard prediction | 5 | On | 200 | Good balance for most cases |
| High-accuracy | 5–10 | On | 300–500 | Use for final candidates |
| Conformational diversity | 10 | On | 200 | Captures structural diversity |
When to use constraints
Pocket constraints are valuable when:
- You know the binding site from crystallography
- Mutagenesis data indicates specific binding residues
- You want to dock a ligand to a specific location
Contact constraints are useful for:
- Incorporating crosslinking mass spectrometry data
- Modeling known protein-protein interfaces
- Forcing specific domain orientations