Key takeaways
- Catalogued protein sequence clusters: 475,217,233 in UniRef100 (UniProt REST API release 2026_01, accessed April 23, 2026)
- Human protein-coding genes: 19,433 in current GENCODE v49
- Distinct translated human products: 129,801 in current GENCODE v49
- Human Proteome Project reference proteome: 19,435 proteins, with 93.6% confidently detected (2025 HUPO HPP report, published 2026)
- Protein molecules in a typical mammalian cell: about 10 billion (2023 review)
As of April 23, 2026, the cleanest single answer is that UniProt's UniRef100 database contains 475,217,233 protein sequence clusters. That number matters because it describes how much sequence space biology has actually catalogued. But there is no single universal protein count: in humans, you can also count 19,433 protein-coding genes, 129,801 distinct translated products, a 19,435-protein Human Proteome Project reference proteome with 93.6% confident detection, or about 10 billion protein molecules in one typical mammalian cell.
There is no single protein count
Protein counts range from 19,433 human protein-coding genes to 475,217,233 UniRef100 sequence clusters because different sources count different biological objects.
If you mean known sequences, the relevant number is a database count such as UniRef100. If you mean human gene products, the relevant numbers come from gene annotation sets such as GENCODE and HPP reference lists. If you mean physical molecules inside cells, the number is much larger, because one protein type can exist in thousands to billions of copies.
That is why articles about protein counts often seem to disagree while all being partly correct.
UniRef currently indexes 475.2 million protein sequence clusters
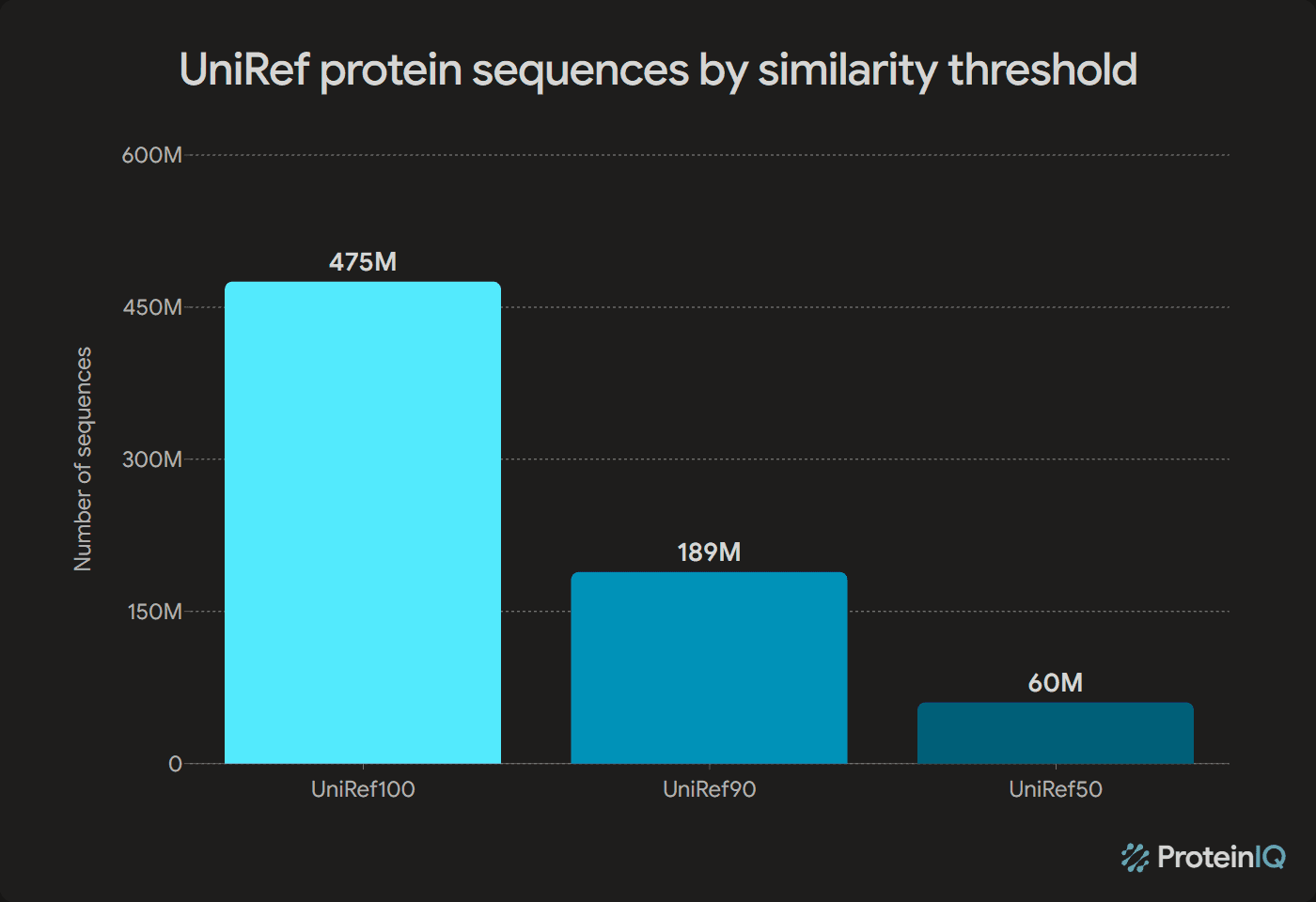
UniProt's UniRef API returned 475,217,233 clusters at 100% identity, 188,848,220 clusters at 90% identity, and 60,315,044 clusters at 50% identity when queried on April 23, 2026.[1]UniRef REST API counts for UniRef100, UniRef90, and UniRef50UniProt · April 23, 2026View source
These are not three different answers to the same question. They are three levels of redundancy reduction. UniRef100 merges exact sequence matches, UniRef90 groups closely related sequences, and UniRef50 compresses the database further into broader protein families.
| UniRef dataset | What it counts | Count |
|---|---|---|
| UniRef100 | Exact-sequence clusters | 475,217,233 |
| UniRef90 | Clusters at 90% sequence identity | 188,848,220 |
| UniRef50 | Clusters at 50% sequence identity | 60,315,044 |
Source: UniProt REST API queries for UniRef100, UniRef90, and UniRef50, accessed April 23, 2026. Response headers reported UniProt release 2026_01.[1]UniRef REST API counts for UniRef100, UniRef90, and UniRef50UniProt · April 23, 2026View source
The human proteome is 19,433 genes, 129,801 translated products, and 93.6% confidently detected in HPP
GENCODE v49 lists 19,433 human protein-coding genes, 211,446 protein-coding transcripts, and 129,801 distinct translations.[2]Human release statistics (v49)GENCODEView source
Those are annotation counts: they describe what the current reference gene set says the human genome can encode. The Human Proteome Project asks a different question: how much of the reference proteome has confident evidence of expression?
The 2025 HUPO Human Proteome Project report describes an HPP reference proteome of 19,435 proteins based on GENCODE v48, UniProtKB 2025_03, Human Protein Atlas 24, MassIVE-KB 2023, and PeptideAtlas 2025-01. It reports that 93.6% of that proteome has been detected.[3]The 2025 Report on the Human Proteome from the HUPO Human Proteome ProjectJournal of Proteome Research · 2026View source Human Protein Atlas summarized the same 2025 report as 19,435 protein-coding genes with 94% confident PE1 detection.[4]The 2025 HUPO HPP report on the human proteomeHuman Protein Atlas · 2026View source
| Human proteome level | Count | What it means |
|---|---|---|
| Protein-coding genes | 19,433 | Genes annotated as protein-coding in GENCODE v49 |
| Protein-coding transcripts | 211,446 | Transcript isoforms annotated as protein-coding |
| Distinct translations | 129,801 | Distinct translated protein products in GENCODE v49 |
| HPP reference proteome | 19,435 proteins | 2025 HUPO HPP target list based on GENCODE v48 plus integrated protein resources |
| Confidently detected HPP proteins | 93.6% | Share of the 2025 HPP reference proteome detected with confident expression evidence |
This separation matters. A gene count is not a protein count, an annotated translation count is not the same thing as a protein that has been directly observed in experiments, and the current GENCODE v49 gene count does not have to match the 2025 HPP target list exactly because the HPP report used GENCODE v48 plus additional protein resources.
A typical mammalian cell contains about 10 billion protein molecules
A 2023 review on protein counting and single-molecule proteomics says a typical mammalian cell of roughly 3,000 um3 contains about 10,000,000,000 protein molecules, with a typical density of about 3 million protein molecules per cubic micrometer.[5]Sampling the proteome by emerging single-molecule and mass spectrometry methodsNature Methods · 2023View source
That is a molecule count, not a count of unique protein types. Deep mass-spectrometry studies can identify around 10,411 protein groups in a 30-minute human cell-line proteomics run, which shows the gap between counting protein copies and counting protein species.[6]The One Hour Human ProteomePubMed · 2024View source
For a practical mental model:
| Cell-level measure | Typical value | Source |
|---|---|---|
| Total protein molecules in a mammalian cell | ~10 billion | 2023 review |
| Protein density | ~3 million molecules per um^3 | 2023 review |
| Protein groups identified in a fast deep human proteome run | 10,411 | 2024 proteomics study |
Protein sequence space is far larger than biology has sampled
For a protein just 100 amino acids long, the number of possible sequences is 20100 because each position can hold one of 20 standard amino acids.
That theoretical space is so large that the 475.2 million catalogued UniRef100 clusters represent only a tiny, biologically explored corner of what chemistry allows. This is one reason protein engineering and protein design still have so much open search space.
Methodology
This article uses four different counting frames, and they should not be merged into a single headline number.
- Known sequence clusters come from live UniProt UniRef REST API queries run on April 23, 2026. The counts are read from the
X-Total-Resultsresponse header. - Human annotated genes, transcripts, and translations come from the current GENCODE human statistics page, which reports release v49.[2]Human release statistics (v49)GENCODEView source
- Human protein detection status comes from the 2025 HUPO Human Proteome Project report. Its HPP reference proteome count is not substituted for the current GENCODE v49 gene count because it was built from GENCODE v48 plus integrated proteomics resources.[3]The 2025 Report on the Human Proteome from the HUPO Human Proteome ProjectJournal of Proteome Research · 2026View source
- Protein molecules per cell refer to molecules, not unique protein types, and come from a review that summarizes cell volume and molecule-density estimates for a typical mammalian cell.[5]Sampling the proteome by emerging single-molecule and mass spectrometry methodsNature Methods · 2023View source
The theoretical 20100 sequence-space figure is a simple combinatoric calculation, not a database count.
Sources▼
- UniRef REST API counts for UniRef100, UniRef90, and UniRef50 UniProt · April 23, 2026. https://rest.uniprot.org/uniref/search?size=1&query=identity%3A1.0
- Human release statistics (v49) GENCODE. https://www.gencodegenes.org/human/stats.html
- The 2025 Report on the Human Proteome from the HUPO Human Proteome Project Journal of Proteome Research · 2026. https://pubs.acs.org/doi/full/10.1021/acs.jproteome.5c00759
- The 2025 HUPO HPP report on the human proteome Human Protein Atlas · 2026. https://www.proteinatlas.org/news/2026-02-20/the-2025-hupo-hpp-report-on-the-human-proteome
- Sampling the proteome by emerging single-molecule and mass spectrometry methods Nature Methods · 2023. https://www.nature.com/articles/s41592-023-01802-5
- The One Hour Human Proteome PubMed · 2024. https://pubmed.ncbi.nlm.nih.gov/38579929/