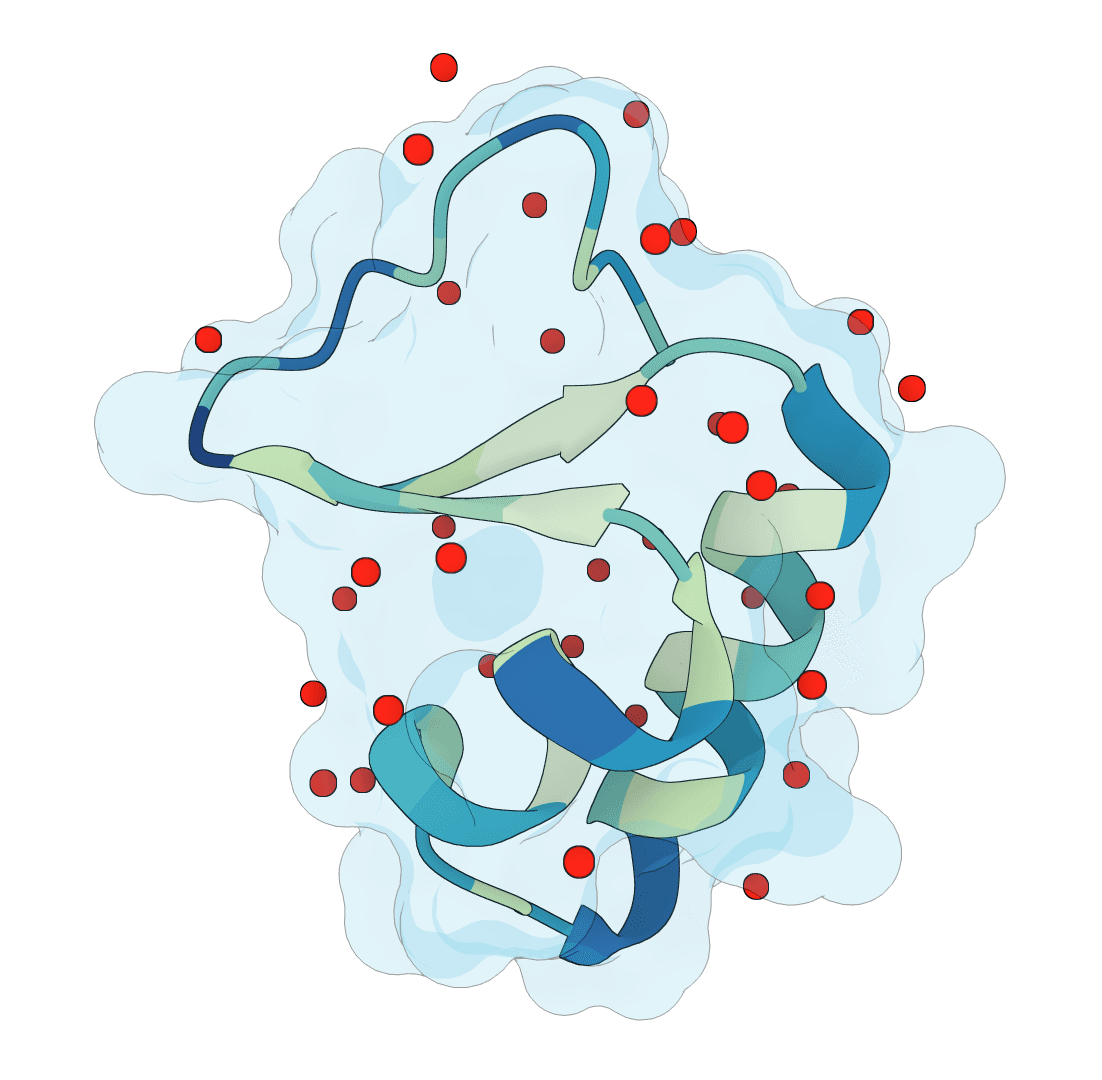
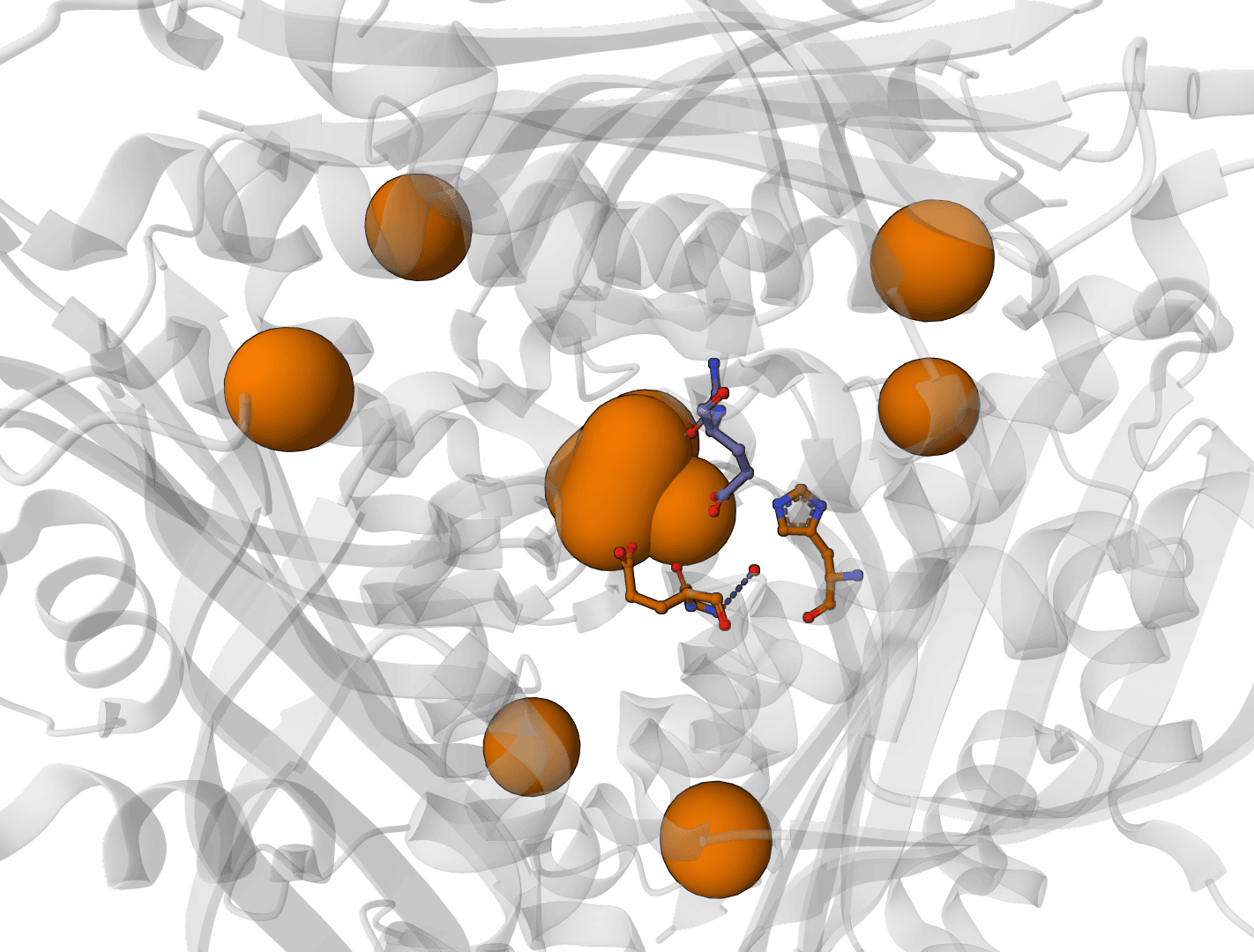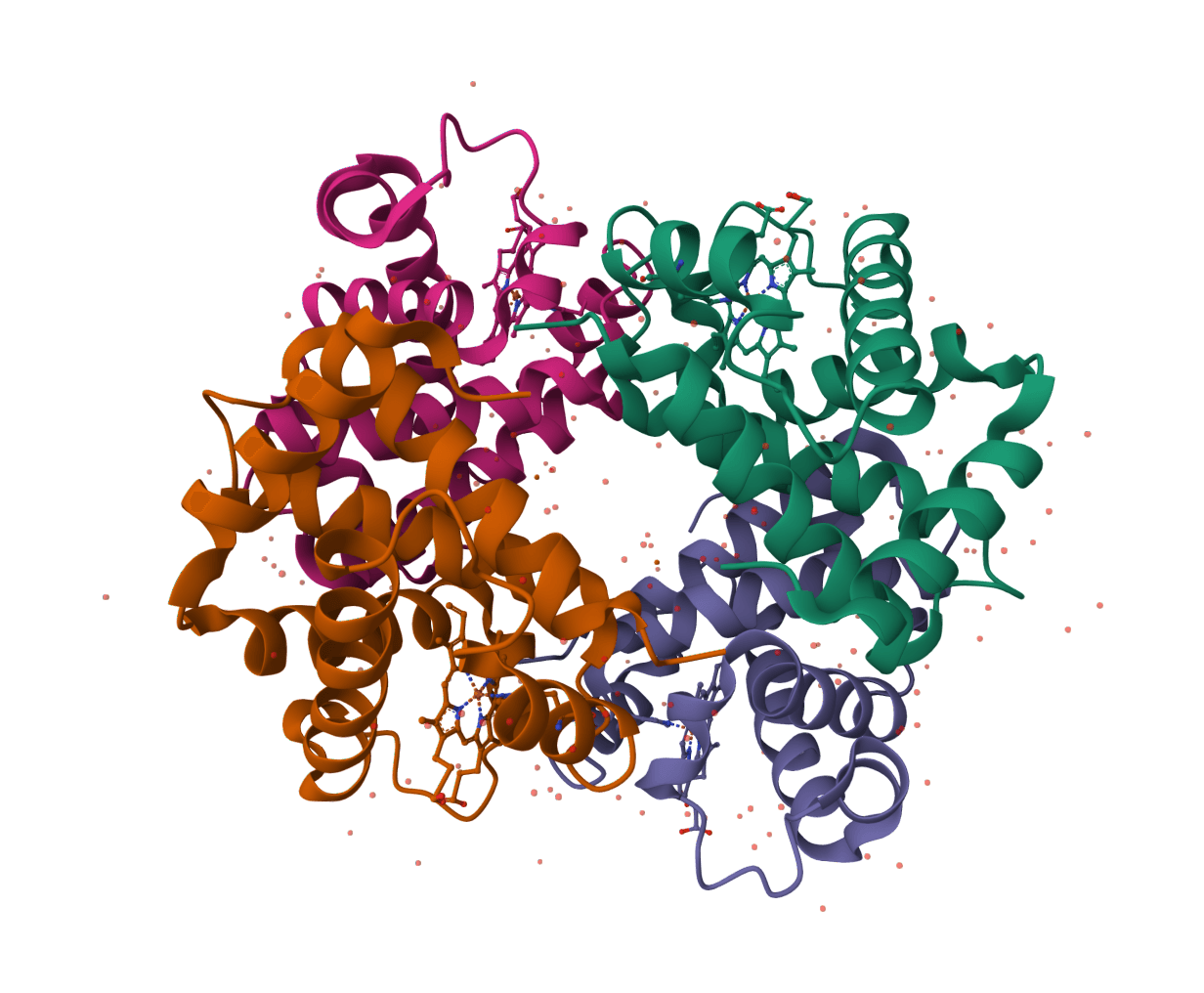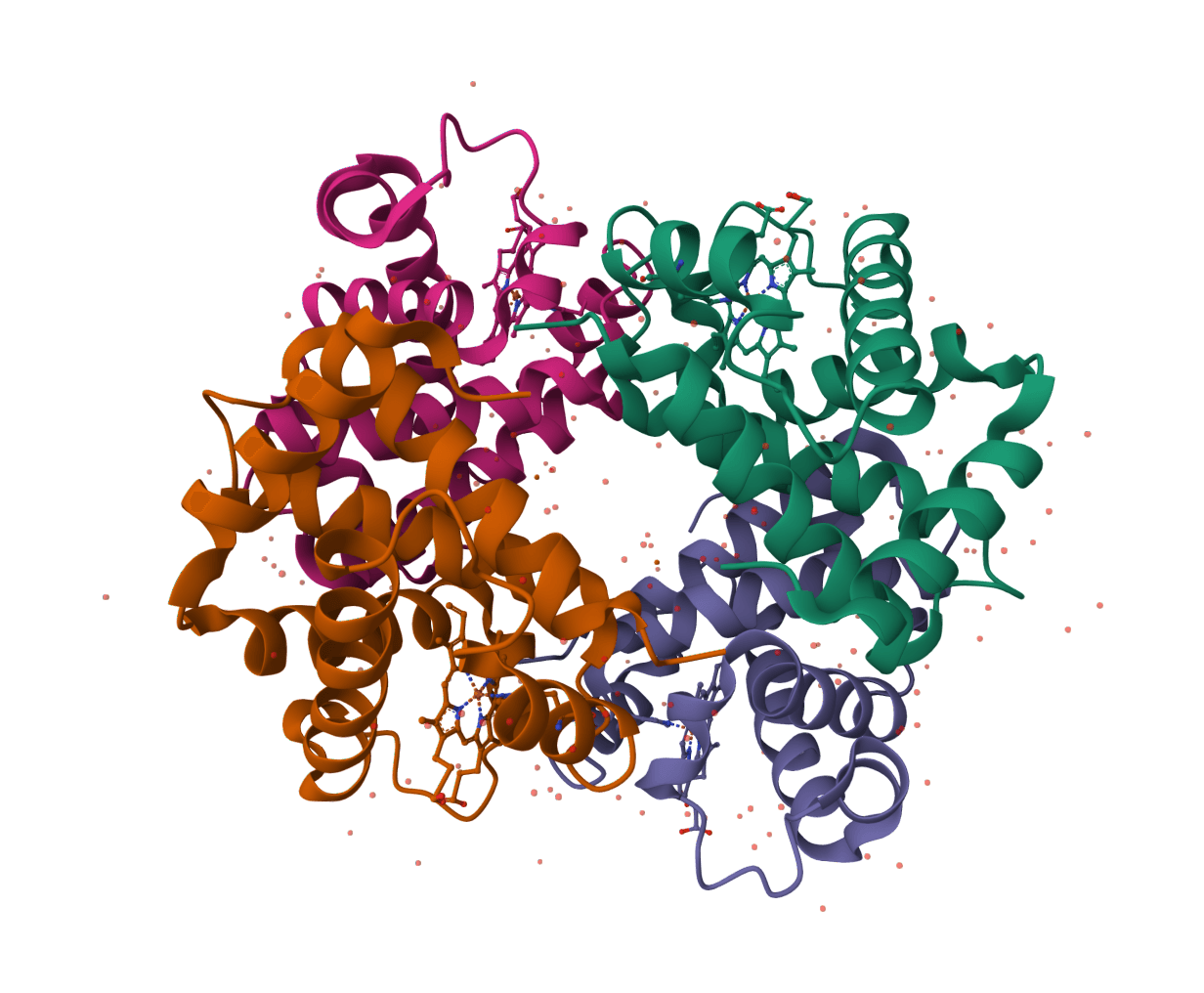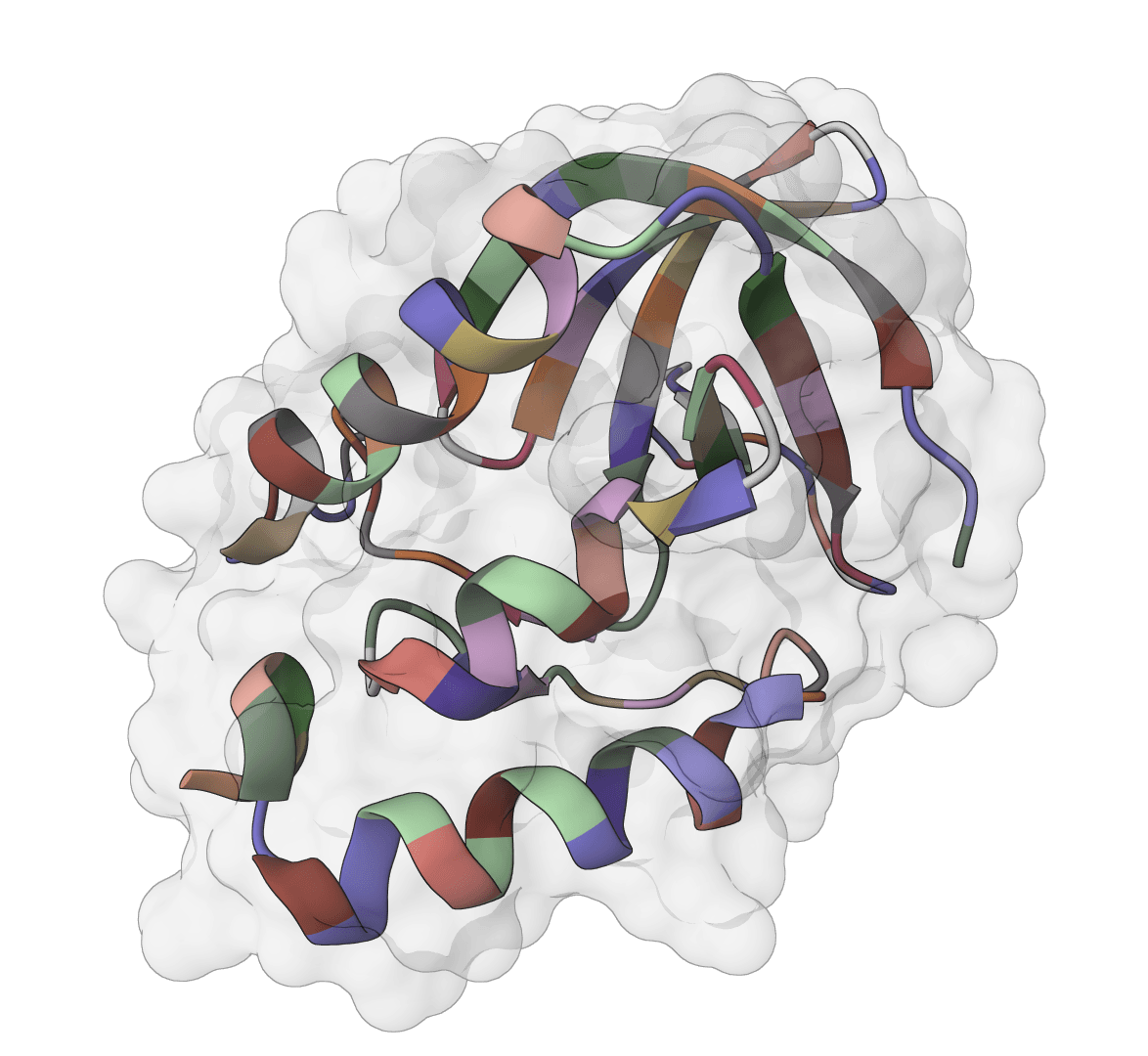What is SuperWater?
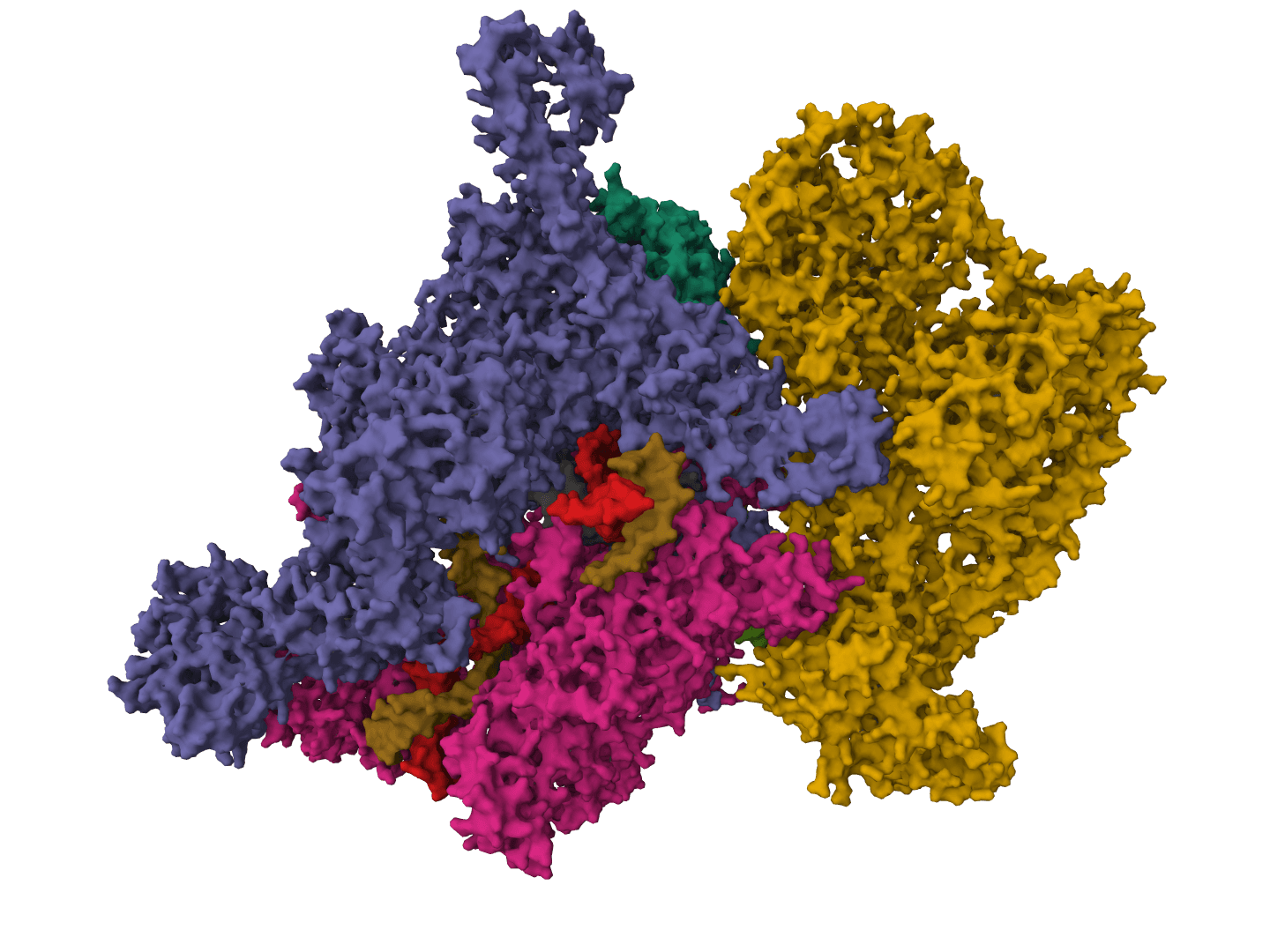
SuperWater predicts ordered water molecule positions around protein structures. It uses a generative model to sample candidate hydration sites, then filters and clusters those candidates to recover likely crystallographic and interface waters from a single structure.
Published in Communications Chemistry in December 2025, SuperWater was introduced as a score-based diffusion framework with equivariant graph neural networks and ESM-derived features. In the reported benchmarks, it outperformed HydraProt and GalaxyWater-CNN across much of the precision-coverage range for protein surface waters as well as protein-protein and protein-ligand interface waters.
Applications
- Structure interpretation: Recovering ordered hydration sites that help stabilize local folds and hydrogen-bond networks
- Binding-site analysis: Highlighting bridging waters near protein-ligand interfaces that may influence affinity and selectivity
- Protein-protein interfaces: Identifying waters that mediate contacts between chains
- Model refinement: Adding plausible solvent positions before visualization, inspection, or downstream structural analysis
How to use SuperWater online
ProteinIQ provides browser-based access to SuperWater on hosted compute, so hydration-site prediction can be run from an uploaded structure or an RCSB entry without local setup.
Settings
Output
ProteinIQ returns a 3D viewer, a tabular summary, and downloadable result files.
Output columns
How does SuperWater work?
SuperWater does not score a fixed 3D voxel grid the way earlier hydration predictors do. Instead, it learns the gradient of the water-position distribution around a protein and uses that learned score field to refine randomly initialized water coordinates through reverse diffusion.
The architecture reported in the paper combines a score-based diffusion model with equivariant graph neural networks, which preserves geometric consistency under rotation. The published method also incorporates ESM features to encode sequence-derived context around residues. After sampling, a separate confidence model removes low-probability positions, and a clustering step consolidates nearby candidates into final hydration sites.
The benchmark protocol described in the paper sampled an initial number of candidates proportional to protein length, then traced different precision-coverage tradeoffs by varying the internal confidence threshold (cap). On an independent test set of 1,709 crystal structures, SuperWater defined the best overall precision-coverage frontier among the compared methods across many operating points.
Interpreting results
SuperWater predictions represent likely ordered water sites, not every transient solvent molecule around the protein. High-confidence predictions are more likely to correspond to persistent hydration sites that recur in experimental structures or stabilize interfaces.
The paper reports mean absolute deviation of approximately 0.3 ± 0.06 Å at cap = 0.5 for matched predictions, which indicates that true-positive sites can be placed with sub-angstrom accuracy. That figure should be interpreted as benchmarked localization accuracy against crystallographic waters, not as a guarantee for every structure.
Limitations
SuperWater predicts positions from a static protein structure. It does not explicitly model long-timescale solvent dynamics, alternate conformations, protonation-state uncertainty, or experimental conditions such as crystal packing, buffer composition, or ligand occupancy.
Like other hydration-site predictors, it is biased toward ordered waters that are recoverable from structural datasets. Disordered, low-occupancy, or rapidly exchanging solvent molecules may be absent from the output even when they are biologically relevant.
Prediction counts also depend directly on Water ratio and Confidence cutoff. A denser sampling strategy can improve recall, but it does not by itself validate the biological importance of every returned site.
ProteinIQ accepts large structures by routing them across multiple GPU tiers, but there is still a hard ceiling for very large residue count × water ratio combinations. If a submission exceeds that supported range, lower Water ratio or use a smaller structure.