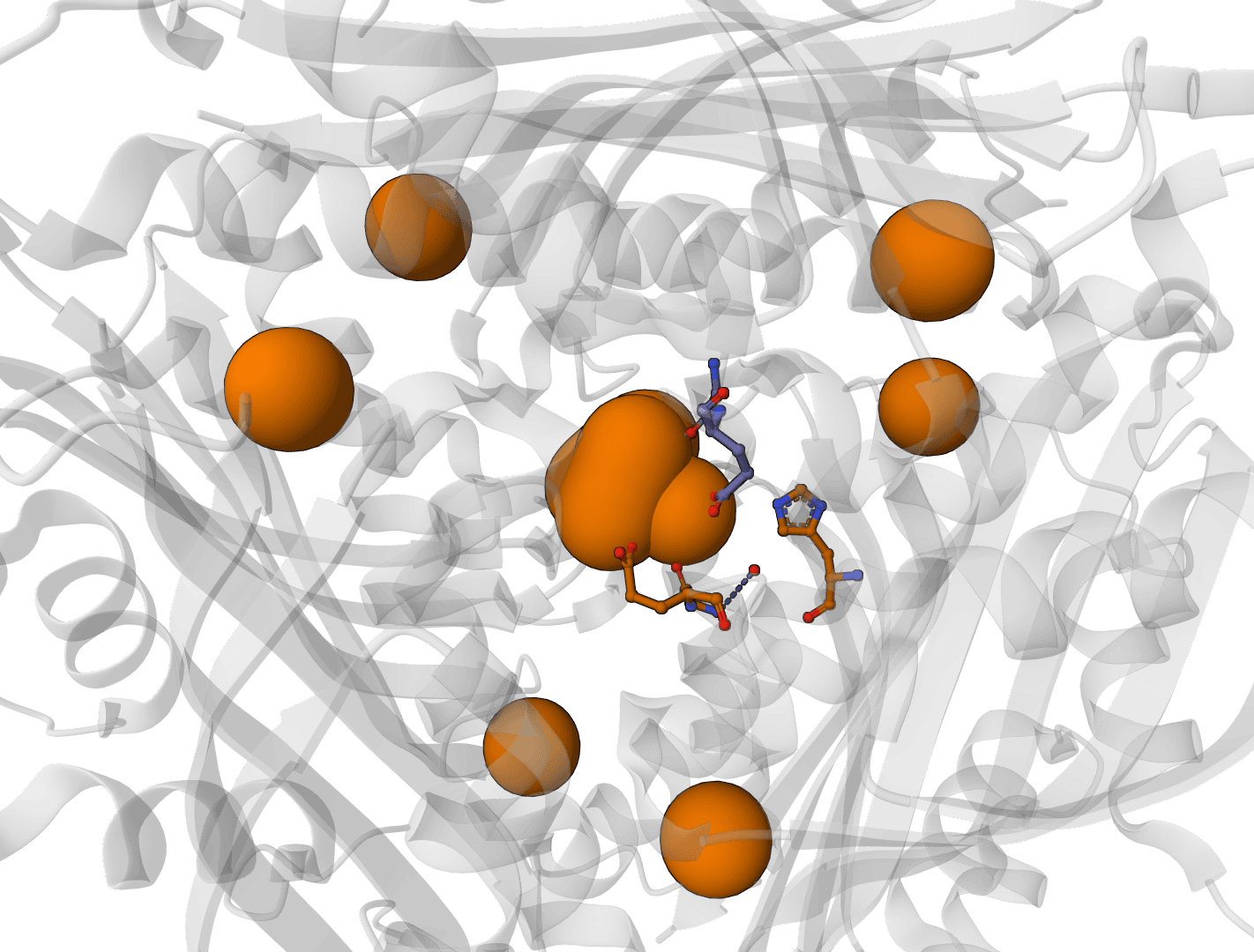Related tools
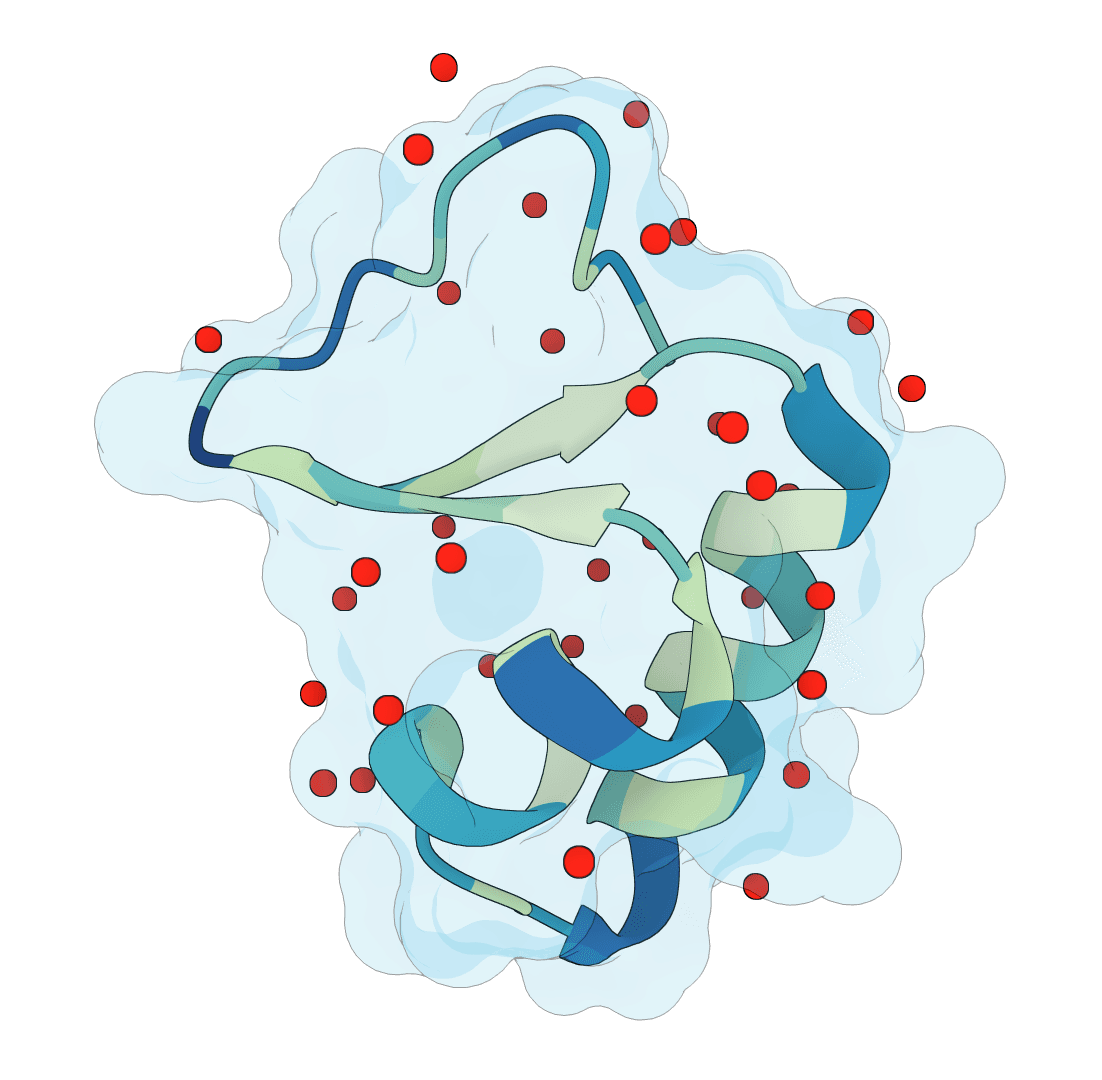
SuperWater
Predict protein hydration sites from a structure using a diffusion model with ESM features and a confidence-filtering head.
Aggrescan3D
Faithful static-mode Aggrescan3D wrapper for per-residue aggregation propensity analysis from a single protein structure.
PROPKA 3
Predict pKa values of ionizable groups in proteins and protein-ligand complexes from 3D structure. PROPKA calculates environment-driven pKa shifts for standard ionizable residues, terminal groups, and supported ligand atom types.
ThermoMPNN
Predict protein thermostability changes (ΔΔG) for point mutations using a graph neural network. Enables computational saturation mutagenesis screening to identify stabilizing mutations.
MolProbity
Validate protein structure quality with all-atom contact analysis, Ramachandran plots, rotamer assessment, and geometry checks.
Protein stability
Predict protein stability using validated BioPython methods: Instability Index, Aliphatic Index, GRAVY, flexibility analysis, and charge distribution
ScanNet
Geometric deep learning model for predicting protein binding sites directly from 3D structure. Identifies where proteins interact with other proteins, antibodies, or disordered proteins with high accuracy, including for novel protein folds.
DockQ
Assess docking model quality by comparing predicted complexes against native references. DockQ v2.1.3 supports protein, nucleic-acid, and supported small-molecule interfaces with faithful upstream metrics.
DSSP
Assign protein secondary structure using the DSSP algorithm. The gold standard for hydrogen bond-based structure assignment from coordinates.
IPSAE
Scoring function for interprotein interactions in AlphaFold2, AlphaFold3 and Boltz predictions. Calculates ipSAE, ipTM, pDockQ, pDockQ2, and LIS scores to assess protein-protein interface quality.
What is AllMetal3D?
AllMetal3D predicts metal binding sites in protein structures, including where a site is likely to occur, the most likely upstream identity class, and the coordination geometry. Its identity classifier reports the same classes exposed by the upstream project: Alkali, MG, CA, ZN, NonZNTM, and NoMetal. A companion model, Water3D, predicts likely water binding positions in the same framework.
Developed by Simon Duerr and Ursula Roethlisberger at EPFL's Laboratory of Computational Chemistry and Biochemistry, AllMetal3D addresses a persistent blind spot in structure prediction: deposited PDB structures frequently have missing or misassigned metal ions, and apo structures lack metals entirely. AllMetal3D benchmarks favorably against MetalSiteHunter, AlphaFold3, MIC, MIB2, and MetalHawk.
How to use AllMetal3D online
ProteinIQ runs AllMetal3D and Water3D on cloud GPU infrastructure—no software installation, no Python environment.
Input
| Input | Details |
|---|---|
Protein Structure | PDB, ENT, CIF, or mmCIF file, or a RCSB PDB ID. The structure must contain protein atoms. Maximum file size 50 MB. |
Settings
| Setting | Description |
|---|---|
Models | AllMetal3D + Water3D (default) runs both models. Selecting one model reduces runtime. |
Prediction mode | Fast (default) subsamples residues for speed. All residues scans every residue and is more thorough. Site-focused restricts analysis to a sphere around specified residues. |
Central residue | Required for site mode only. One or more residue numbers separated by spaces (e.g., 101 203). |
Site radius | Radius in Ångströms around the central residues in site mode. Range 4–50 Å, default 8 Å. |
Clustering threshold | Agglomerative clustering cutoff in Ångströms. Lower values split nearby predictions into separate sites; higher values merge them. Range 1–10 Å, default 7 Å. |
Probability threshold | Minimum confidence for a site to appear in results. Raise to see only high-confidence predictions. Range 0.05–1.0, default 0.25. |
Batch size | Inference batch size. Lower values (e.g., 10) reduce GPU memory usage on very large structures. Default 50. |
Output
Results for AllMetal3D are tabulated by site:
| Column | Description |
|---|---|
Structure | Source structure identifier |
Site | Sequential site index |
X, Y, Z | Predicted metal coordinates in Ångströms |
Location confidence | Model confidence in the predicted position (%) |
Predicted identity | Most likely upstream identity class (Alkali, MG, CA, ZN, NonZNTM, or NoMetal) |
Identity confidence | Confidence in the metal identity classification (%) |
Predicted geometry | Coordination geometry (e.g., tetrahedral, octahedral) |
Geometry confidence | Confidence in the geometry classification (%) |
The detailed table also includes raw per-class probability columns for both identity and geometry, so you can inspect the full upstream distribution instead of only the top-ranked class.
Downloadable outputs include the predicted PDB (with metal coordinates appended) and CUBE-format electron density files for visualization.
How AllMetal3D works
AllMetal3D uses a fully convolutional 3D CNN trained on metal-containing structures from the PDB. The network processes a volumetric representation of the local protein environment centered on candidate positions. The architecture consists of five convolutional layers with 8, 60, 100, 80, and 30 channels, with leaky ReLU activation (slope 0.2) and max-pooling after the second layer. Processed features condense into a 1,280-dimensional fingerprint that feeds into separate fully-connected heads for identity and geometry classification.
Prediction follows a two-stage pipeline. AllMetal3Dloc first identifies candidate positions by scanning the structure and scoring each position for the likelihood of containing any metal. Positions above the probability threshold are then passed to the classification network, which outputs the most probable metal identity and coordination geometry.
Water3D applies the same 3D CNN framework trained specifically on crystallographic water positions, identifying ordered water molecules that are consistently present across multiple crystal structures of related proteins.
Interpreting results
Metal identity confidence
Identity classification accuracy varies substantially by metal type. Zinc sites are predicted most reliably given the abundance of Zn-containing structures in training data. Magnesium and sodium are more challenging due to their broad and irregular binding motifs.
| Confidence | Interpretation |
|---|---|
| >70% | High-confidence assignment—treat as reliable |
| 40–70% | Moderate—consider the predicted identity likely but not certain |
| <40% | Low—the model sees a metal-like environment but cannot distinguish identity well |
Coordination geometry
The geometry classifier performs well for tetrahedral and octahedral arrangements, which dominate in natural proteins. Other geometries (square planar, trigonal bipyramidal, etc.) are classified less accurately and should be treated with caution.
Water3D sites
Water predictions reflect ordered, structurally conserved water positions. Loosely bound or disordered waters—which appear at low occupancy in crystal structures—will generally fall below the probability threshold.
Limitations
Metal selectivity depends partly on cellular context (compartmentalization, metal availability during protein synthesis) that cannot be inferred from structure alone. A site predicted as Mg²⁺ may be occupied by other divalent ions in different cellular environments.
Median positional error for metal site location is approximately 0.5 Å. This is sufficient for identifying which residues coordinate the metal, but marginal for distinguishing subtle geometry differences. Vacancy prediction—identifying sites that are empty in a given structure but could be occupied—was not reliable and is not reported.
Performance on Mg²⁺ and Na⁺/K⁺ is meaningfully lower than for transition metals, reflecting sparser training data and the tendency of these ions to adopt irregular or water-mediated coordination environments.