Related tools
DockQ
Assess docking model quality by comparing predicted complexes against native references. DockQ v2.1.3 supports protein, nucleic-acid, and supported small-molecule interfaces with faithful upstream metrics.
MolProbity
Validate protein structure quality with all-atom contact analysis, Ramachandran plots, rotamer assessment, and geometry checks.
PoseBusters
PoseBusters validates generated or docked molecular poses with chemically and structurally grounded quality checks for molecular geometry, intermolecular interactions, and optional reference-pose agreement.
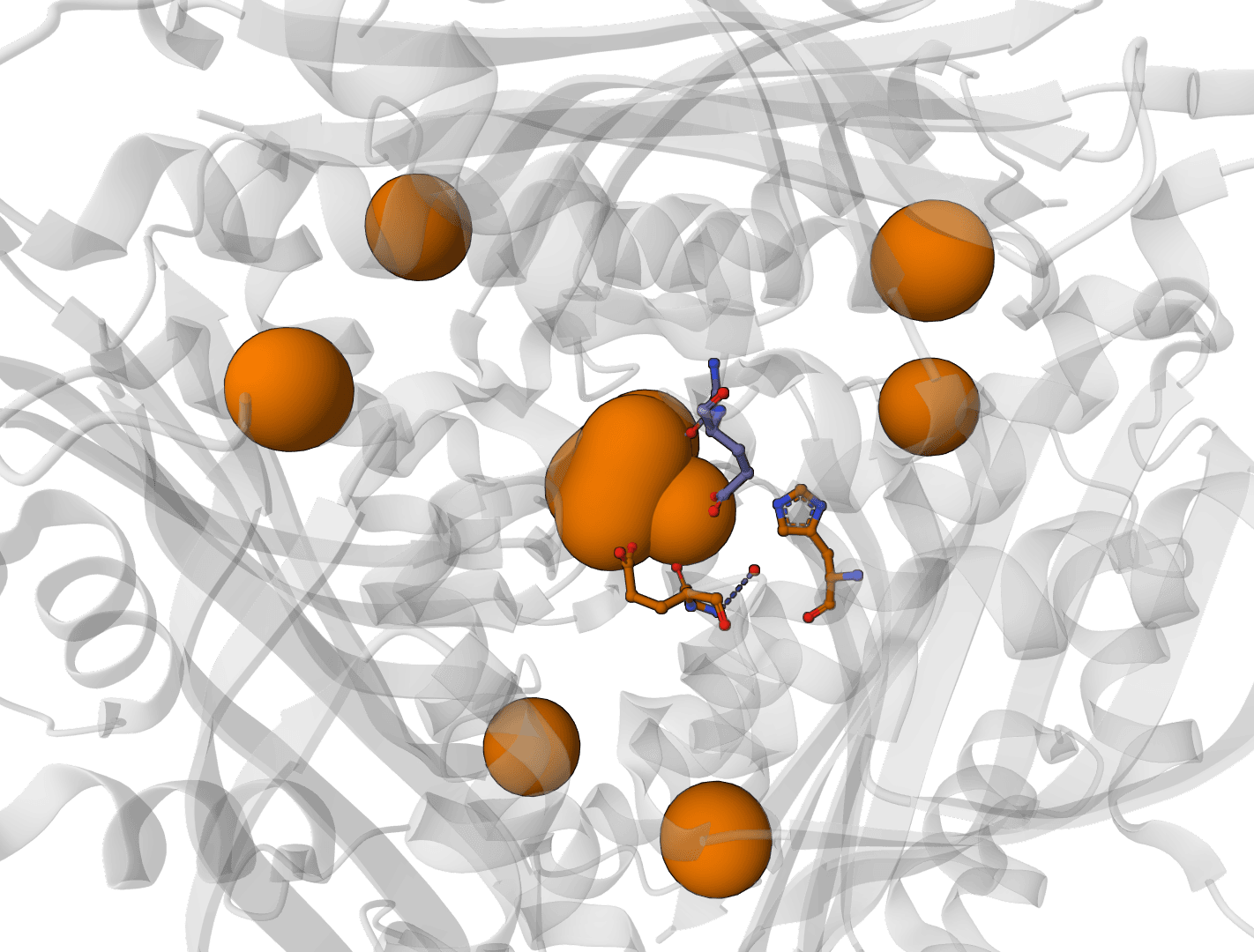
AllMetal3D
Predict metal and water binding sites in protein structures using 3D convolutional neural networks (AllMetal3D + Water3D).
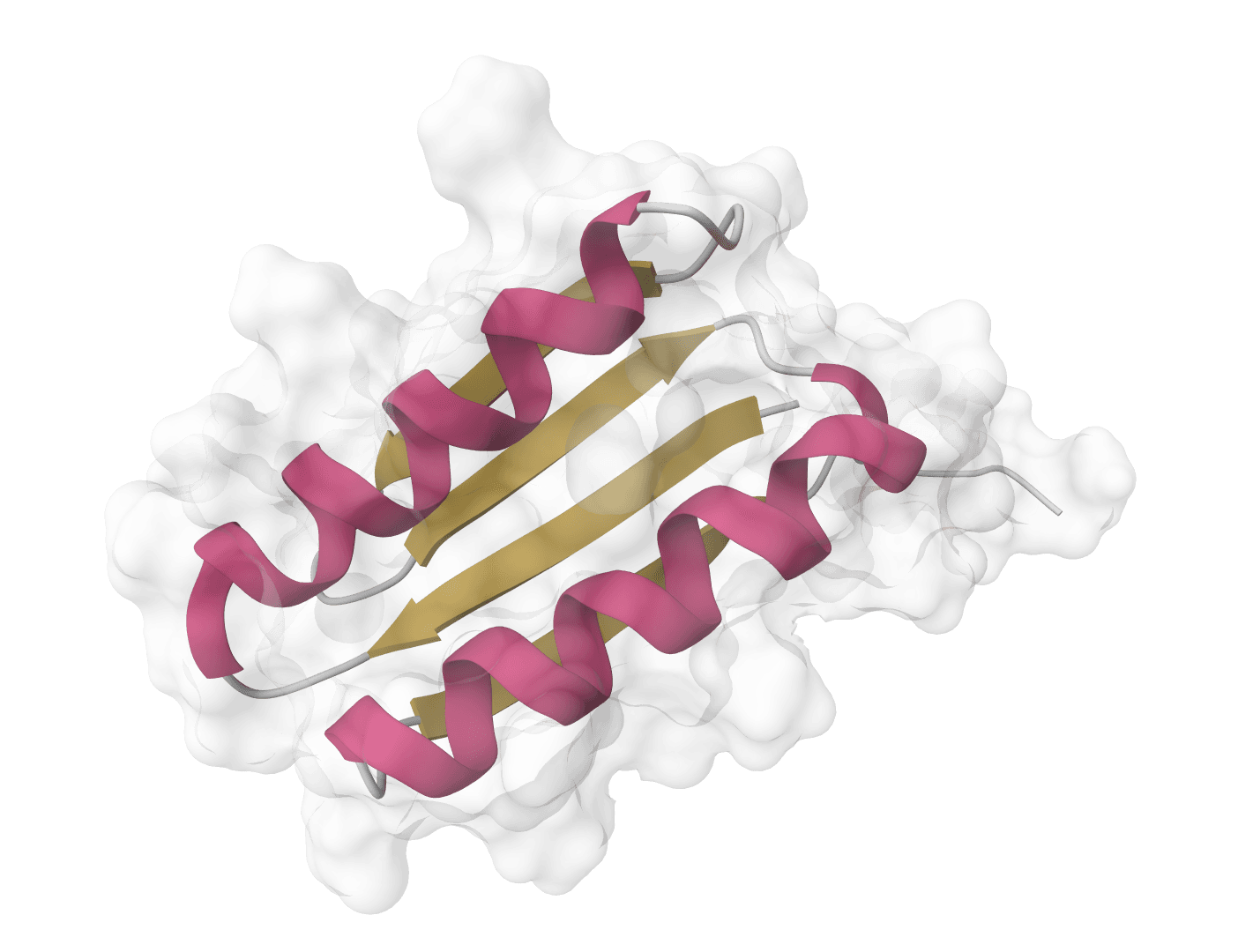
DSSP
Assign protein secondary structure using the DSSP algorithm. The gold standard for hydrogen bond-based structure assignment from coordinates.

Radius of gyration
Calculate the radius of gyration (Rg) for protein structures from PDB files. Supports multiple chains and atom selection options.
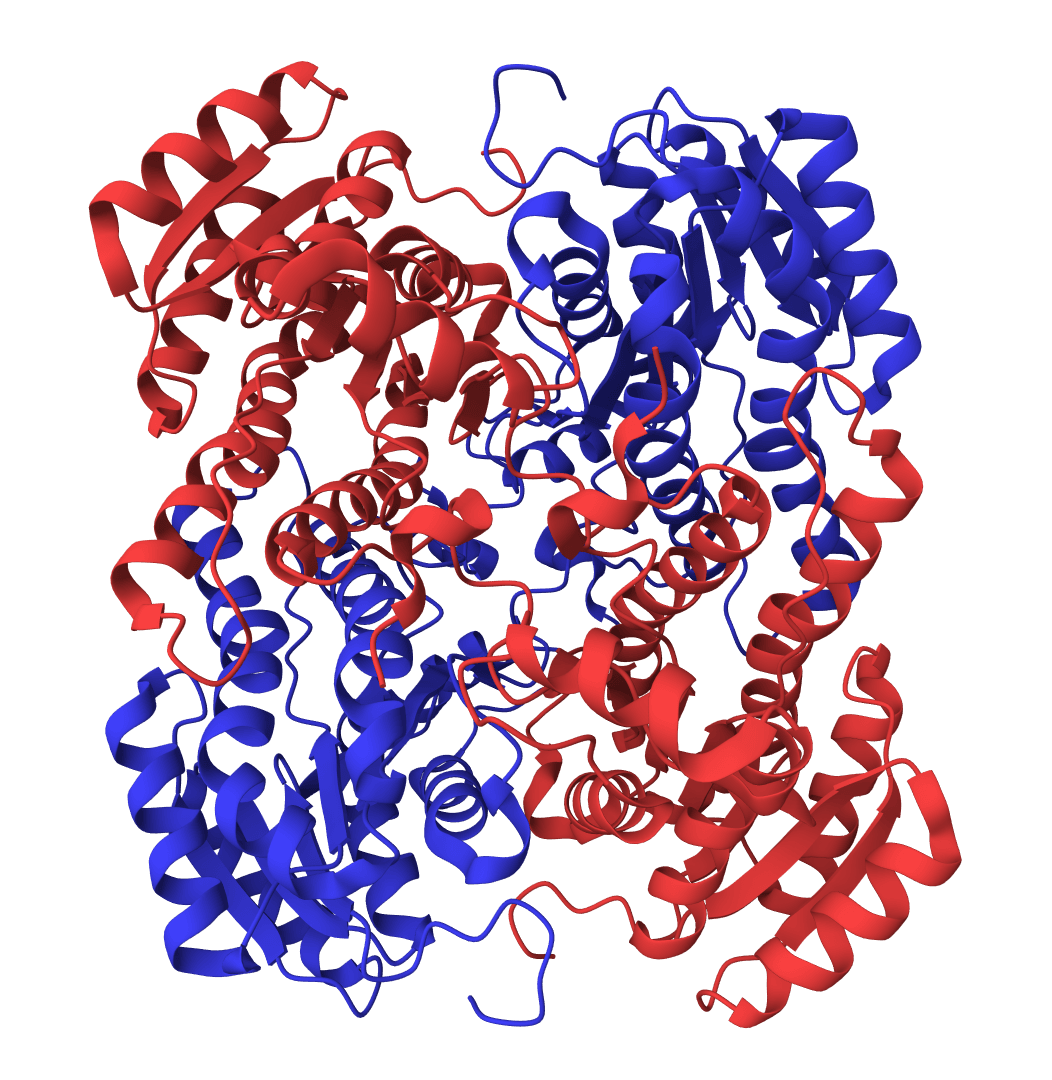
RMSD calculator
Calculate Root Mean Square Deviation (RMSD) between protein structures. Compare a reference PDB against multiple structures with automatic Kabsch alignment.

SASA calculator
Calculate Solvent Accessible Surface Area (SASA) for protein structures using the Shrake-Rupley algorithm.
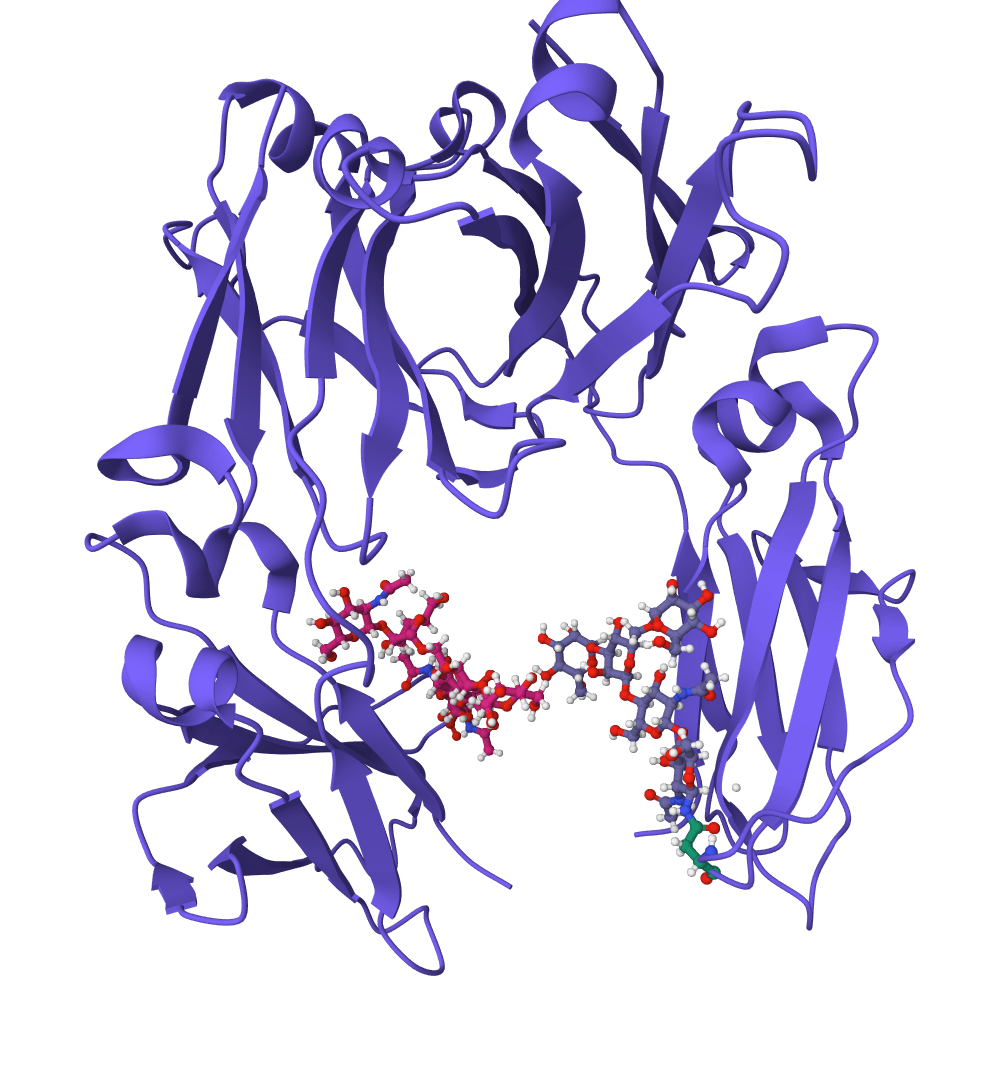
ScanNet
Geometric deep learning model for predicting protein binding sites directly from 3D structure. Identifies where proteins interact with other proteins, antibodies, or disordered proteins with high accuracy, including for novel protein folds.
SuperWater
Predict protein hydration sites from a structure using a diffusion model with ESM features and a confidence-filtering head.
What is IPSAE?
ipSAE (interaction prediction Score from Aligned Errors) is a replacement for AlphaFold's ipTM score, developed by Roland Dunbrack Jr. at the Fox Chase Cancer Center. It addresses a fundamental flaw in how AlphaFold evaluates protein-protein interactions: ipTM averages predicted alignment quality across all residue pairs between chains, including disordered regions and non-interacting domains that drag the score down even when the actual interface is well-predicted.
The problem becomes acute with full-length protein sequences. A kinase dimer predicted with high confidence at its interface can receive a mediocre ipTM simply because both chains contain long disordered tails. Trimming those tails raises the ipTM score despite the structure prediction being identical — a clear sign the metric is measuring the wrong thing.
ipSAE fixes this by filtering out residue pairs with high predicted aligned error (PAE) before scoring, and adjusting the length normalization to reflect only high-confidence residues. In benchmarks on 40 heterodimers with paired decoys, ipSAE cleanly separates true from false interactions even with full-length UniProt sequences, where ipTM struggles.
Beyond ipSAE itself, the tool also computes ipTM, pDockQ, pDockQ2, and LIS (Local Interaction Score) — giving a comprehensive view of interface quality from a single run.
How does ipSAE work?
AlphaFold's ipTM is derived from the TM-score formula:
where scales with protein length. The ipTM applies this across chains, averaging over all interchain residue pairs regardless of prediction quality.
ipSAE modifies this in three ways:
-
PAE filtering: Only residue pairs where the predicted aligned error falls below a cutoff (default 10 Å) contribute to the score. Disordered regions and non-interacting domains are excluded automatically.
-
Adjusted normalization: The parameter is recalculated using only the count of residues that pass the PAE filter, not the total chain length. Short effective lengths () use .
-
Direct PAE values: Instead of AlphaFold's probability distributions over alignment errors, ipSAE uses the actual PAE distances as in the TM formula.
The score is computed asymmetrically — ipSAE(A→B) and ipSAE(B→A) — and the final value is the maximum of the two directions.
Normalization variants
The tool reports three ipSAE variants differing in how is calculated:
| Variant | based on | Best for |
|---|---|---|
ipSAE | Number of residues with PAE below cutoff | General use |
ipSAE (d0=chain) | Full chain length | Comparisons to ipTM |
ipSAE (d0=domain) | Domain-level residue counts | Multi-domain proteins |
How to use IPSAE online
ProteinIQ runs the ipSAE scoring pipeline in the cloud — upload prediction files and get scores back without installing Python dependencies or writing command-line arguments.
Inputs
| Input | Description |
|---|---|
PAE data file | AlphaFold2 JSON, AlphaFold3 *_full_data_*.json, or Boltz NPZ file containing the predicted aligned error matrix. Do not upload AlphaFold3 summary_confidences*.json files. Max 50 MB. |
Structure file | Corresponding PDB or CIF structure file. Also accepts a PDB ID to fetch from RCSB. Max 50 MB. |
Settings
| Setting | Description |
|---|---|
PAE cutoff | Residue pairs with PAE above this value (in Å) are excluded from scoring. Range 1–30, default 10. Lower values are more stringent — a cutoff of 10–15 Å provides optimal discrimination in benchmarks. |
Distance cutoff | Cα–Cα distance threshold (in Å) for identifying interface contacts. Range 5–30, default 15. |
Output columns
| Column | Description |
|---|---|
Chain Pair | The two chains being scored (e.g., A–B). |
Type | Interaction type (heterodimer, homodimer, etc.). |
ipSAE | Primary ipSAE score. Values near 1.0 indicate a confidently predicted interface; near 0 indicates no interaction. |
ipSAE (d0=chain) | ipSAE with normalized by full chain length. |
ipSAE (d0=domain) | ipSAE with normalized by domain residue count. |
ipTM | AlphaFold's original interface predicted TM-score for comparison. |
pDockQ | Interface quality estimate calibrated against DockQ scores. |
pDockQ2 | Improved pDockQ using per-residue contact analysis. |
LIS | Local Interaction Score — measures interaction quality in the immediate interface region. |
Residues Chain 1/2 | Number of residues in each chain. |
Interpreting results
ipSAE scores
ipSAE ranges from 0 to 1. A high score means AlphaFold confidently predicts the interface with low alignment error. A score near zero means AlphaFold found no confident interchain contacts — strong evidence against a physical interaction.
The key advantage over ipTM is visible in borderline cases. For the RAF1-KRAS kinase interaction with full-length sequences, ipTM reports 0.59 (ambiguous), while ipSAE reports 0.80 (clearly positive). For a false pairing like RAF1-RIPK1, ipTM gives 0.28 while ipSAE returns 0.0.
pDockQ and pDockQ2
pDockQ estimates the DockQ quality of a predicted complex based on interface pLDDT and contact count:
| pDockQ | Expected quality |
|---|---|
| > 0.5 | Acceptable or better |
| 0.23–0.5 | Incorrect or borderline |
| < 0.23 | Likely incorrect |
pDockQ2 refines this with per-residue analysis and tends to be more reliable for complexes with multiple interfaces.
Limitations
- ipSAE scores are only meaningful for predictions from AlphaFold2, AlphaFold3, or Boltz — it requires a PAE matrix, which experimental structures do not produce.
- Like all confidence metrics, ipSAE reflects model certainty, not ground truth. A high score means AlphaFold is confident, not that the interaction necessarily occurs in vivo.
- Optimal PAE cutoff values may vary by application. The default of 10 Å works well in published benchmarks, but screening peptide binders or highly flexible interfaces may benefit from a slightly higher cutoff (12–15 Å).