Related tools
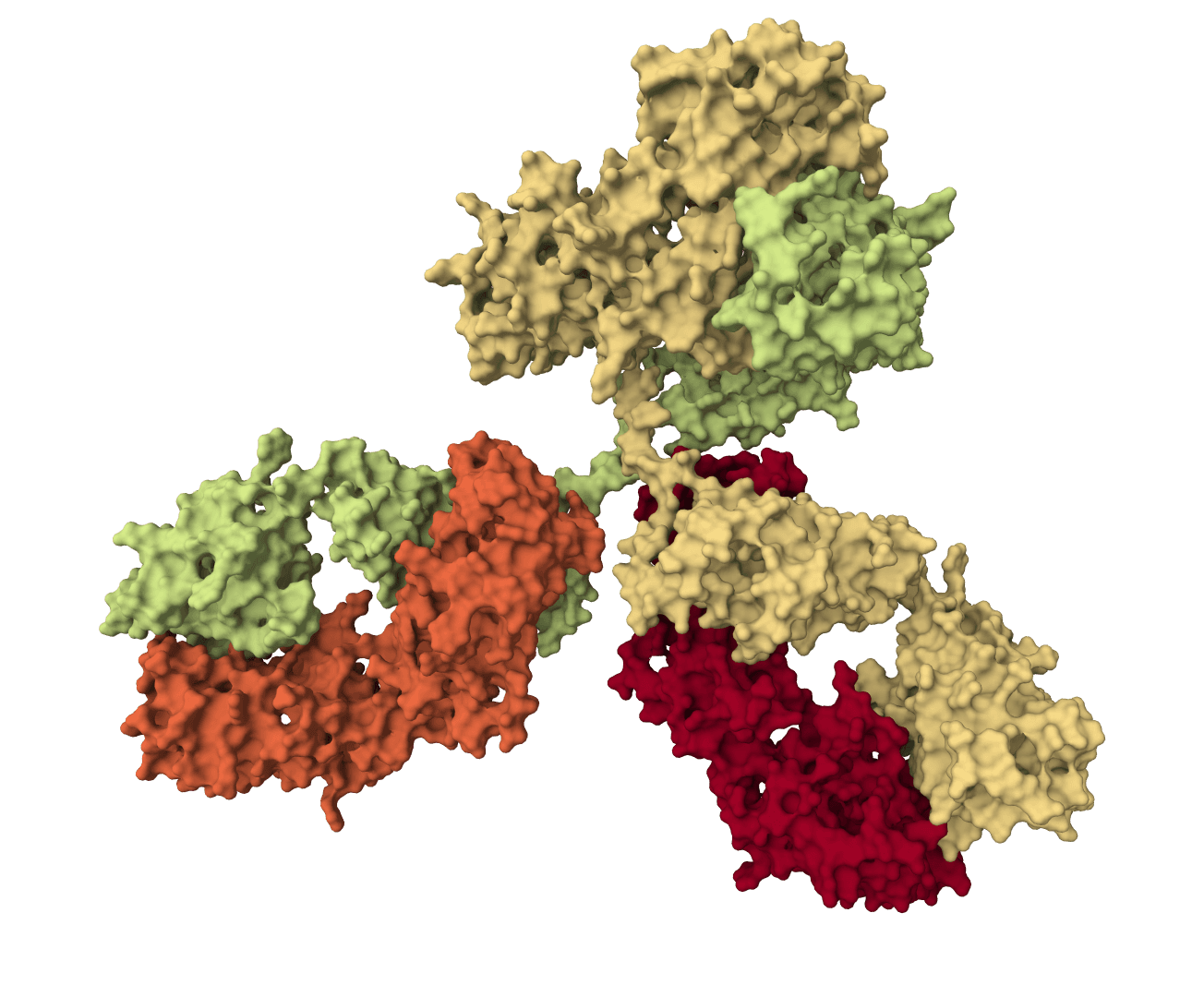
Proteo-R1
Reasoning-guided antibody CDR co-design for antibody-antigen complexes. Proteo-R1 identifies residue-level functional decisions and uses conditional diffusion to generate ranked designed structures with confidence metrics.
BioPhi
Antibody humanization and humanness evaluation platform from Merck. Sapiens mode uses deep learning trained on the Observed Antibody Space (OAS) to humanize antibody sequences, while OASis mode evaluates humanness using 9-mer peptide search against human antibody databases.
BoltzGen
BoltzGen is a state-of-the-art AI model for designing protein and peptide binders against any biomolecular target. Using generative diffusion models, it creates novel binders (proteins, peptides, nanobodies) with nanomolar-level binding affinity.
DiffAb
AI-powered antibody CDR design using equivariant diffusion models. Generates optimized complementarity-determining region (CDR) sequences and structures for antibodies targeting specific antigens. Supports single CDR, multi-CDR co-design, and fixed-backbone sequence design modes.
PepMLM
Design linear peptide binders for target proteins using a target sequence-conditioned masked language model. PepMLM generates peptide sequences optimized to bind specific protein targets based on ESM-2 protein language modeling.
EvoDiff
EvoDiff is a diffusion-based protein sequence generation framework from Microsoft Research. ProteinIQ currently wraps the EvoDiff-Seq OA_DM_38M model for unconditional protein generation, motif scaffolding, and user-sequence inpainting.
ODesign
All-atom generative AI for designing protein binders. Specify target binding sites and generate diverse binding proteins with fine-grained control over interaction parameters.
PocketFlow
PocketFlow is a structure-based molecular generative model that designs novel drug-like molecules within protein binding pockets. It uses autoregressive flow modeling with chemical knowledge to generate 100% chemically valid, highly drug-like compounds.
PocketXMol
PocketXMol is a pocket-interacting generative foundation model for docking, small-molecule design, and peptide design in protein binding pockets.
RFdiffusion
RFdiffusion is a state-of-the-art protein structure generation tool that uses diffusion models to design proteins de novo, create binders, scaffold motifs, and generate symmetric oligomers with atomic precision.
