Related tools
BoltzGen
BoltzGen is a state-of-the-art AI model for designing protein and peptide binders against any biomolecular target. Using generative diffusion models, it creates novel binders (proteins, peptides, nanobodies) with nanomolar-level binding affinity.
RFantibody
Structure-based de novo antibody and nanobody design pipeline combining antibody-tuned RFdiffusion, ProteinMPNN sequence design, and antibody-tuned RoseTTAFold2 filtering.

ProFam
ProFam-1 is a protein family language model for family-conditioned sequence generation. Provide a protein family FASTA/MSA and generate new sequences with model likelihood scores for downstream ranking and screening.
ODesign
All-atom generative AI for designing protein binders. Specify target binding sites and generate diverse binding proteins with fine-grained control over interaction parameters.
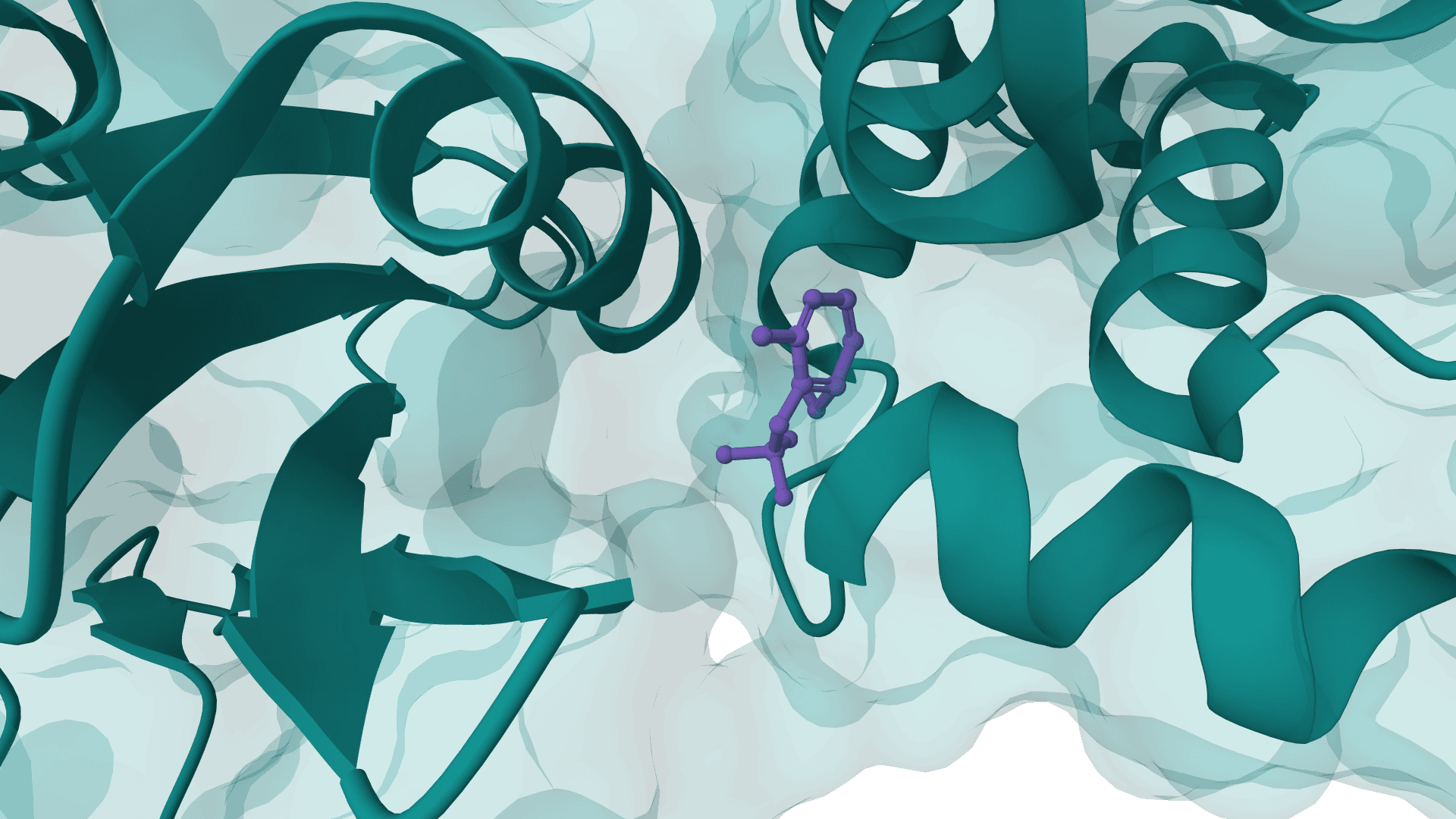
PocketFlow
PocketFlow is a structure-based molecular generative model that designs novel drug-like molecules within protein binding pockets. It uses autoregressive flow modeling with chemical knowledge to generate 100% chemically valid, highly drug-like compounds.
RFdiffusion
RFdiffusion is a state-of-the-art protein structure generation tool that uses diffusion models to design proteins de novo, create binders, scaffold motifs, and generate symmetric oligomers with atomic precision.
EvoDiff
EvoDiff is a diffusion-based protein sequence generation framework from Microsoft Research. ProteinIQ currently wraps the EvoDiff-Seq OA_DM_38M model for unconditional protein generation, motif scaffolding, and user-sequence inpainting.

ProGen2
ProGen2 is Salesforce Research's protein language model suite for prompt-based de novo protein sequence generation. It samples novel amino acid sequences from a plain-text context string using top-p sampling and temperature control.
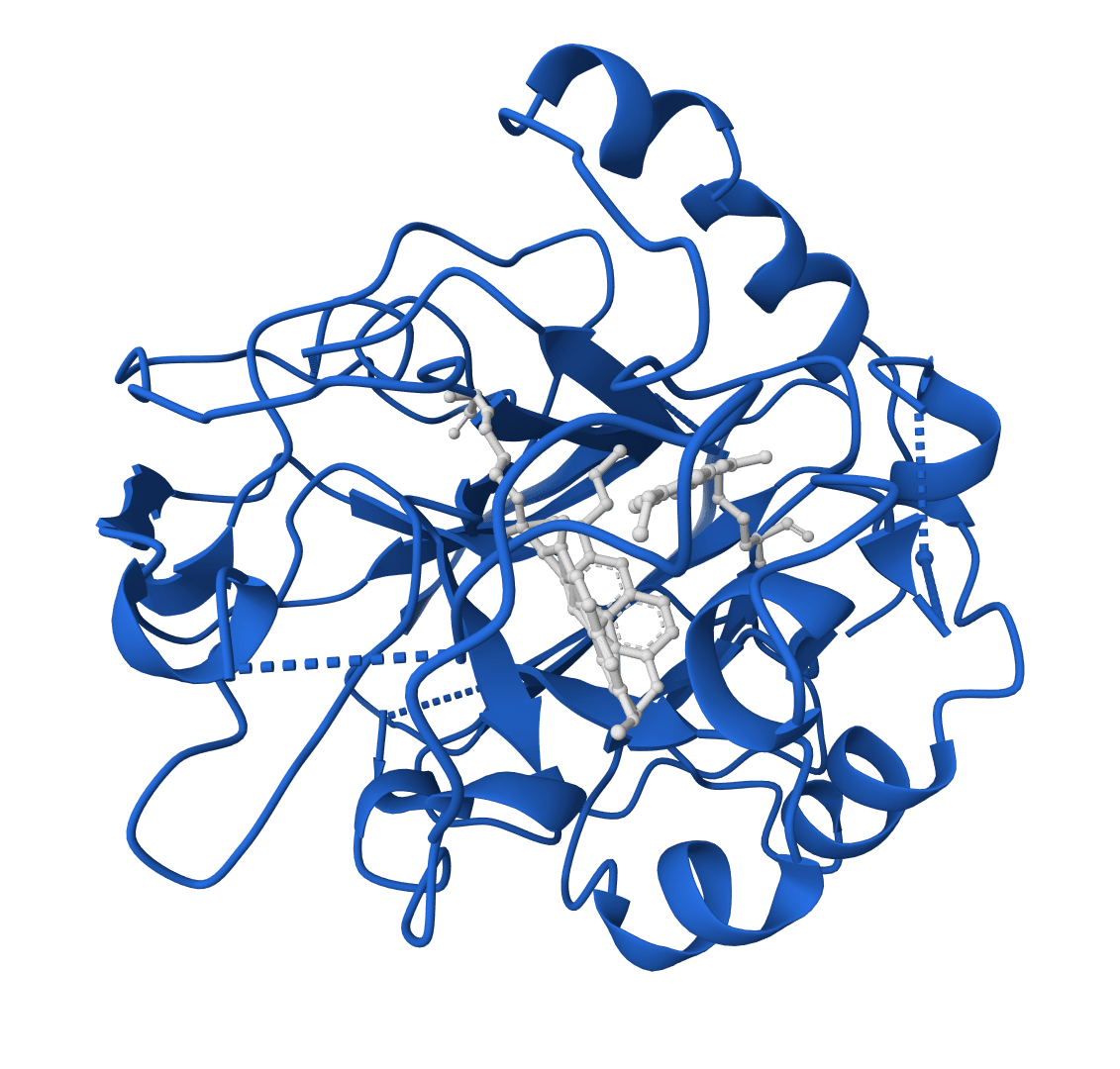
RFdiffusion 2
RFdiffusion2 is an atom-level enzyme active site scaffolding tool that generates protein scaffolds around your input motif. REQUIRES an input PDB structure containing the active site residues to scaffold. For ligand-aware design, ligands must be embedded in the input PDB as HETATM records.
BioPhi
Antibody humanization and humanness evaluation platform from Merck. Sapiens mode uses deep learning trained on the Observed Antibody Space (OAS) to humanize antibody sequences, while OASis mode evaluates humanness using 9-mer peptide search against human antibody databases.
What is PepMLM?
PepMLM is a de novo peptide binder design tool that generates linear peptide sequences predicted to bind specific target proteins. Developed by the Chatterjee Lab and published in Nature Biotechnology in 2025, PepMLM employs a target sequence-conditioned masked language modeling approach built on ESM-2, a state-of-the-art protein language model.
Unlike structure-based design methods that require 3D coordinates of the target protein, PepMLM operates exclusively from amino acid sequences. The model positions peptide binders at the C-terminus of target proteins and reconstructs the binder region through span masking, enabling generative design of candidate binders to any target protein without structural information.
PepMLM was experimentally validated through binding assays and targeted protein degradation studies, demonstrating efficacy against cancer biomarkers, viral phosphoproteins, and Huntington's disease-related proteins.
How to use PepMLM online
ProteinIQ provides a web-based interface for running PepMLM without command-line installation or GPU configuration. Paste or upload target protein sequences, adjust generation parameters, and receive ranked peptide binder candidates with confidence scores.
Inputs
| Input | Description |
|---|---|
Target Protein Sequence(s) | Amino acid sequence of the target protein. Accepts FASTA format, raw text sequences, or UniProt IDs. Multiple targets can be processed in a single job. Maximum 50 sequences per job. |
Settings
Generation parameters
| Setting | Description |
|---|---|
Peptide length | Length of generated binders in amino acids (5–30, default 15). Therapeutic peptides typically range 10–20 residues. Shorter peptides (5–10) offer better cell penetration but may have lower binding affinity. Longer peptides (20–30) may achieve stronger binding but require more complex synthesis. |
Binders per target | Number of candidate peptides to generate for each target sequence (1–20, default 4). Higher values provide greater sequence diversity but increase computation time. |
Top-k sampling | Restricts sampling to the top most probable amino acids at each position (1–10, default 3). Lower values (1–2) produce conservative, high-confidence designs. Higher values (5–10) increase sequence diversity by exploring more of the probability distribution. |
Results
The output presents a ranked table of peptide binder candidates with confidence metrics.
| Column | Description |
|---|---|
Binder | Designed peptide sequence in single-letter amino acid code. Sequences may contain X (any amino acid) at positions where the model assigns similar probabilities to multiple residues. |
Pseudo Perplexity | Language model confidence score for the generated sequence. Lower values indicate higher model confidence. Typical values range from 10–40, with scores below 20 representing high-confidence predictions. |
Input Sequence | Target protein sequence (truncated to 50 characters). Only displayed when processing multiple targets in one job. |
Interpreting pseudo-perplexity
Pseudo-perplexity quantifies how "surprised" the ESM-2 model is by the generated peptide sequence conditioned on the target protein. The metric is calculated exclusively on the masked binder region by measuring reconstruction loss.
- < 15 — Very high confidence, strong sequence likelihood
- 15–25 — High confidence, typical for validated binders
- 25–35 — Moderate confidence, may benefit from experimental screening
- > 35 — Lower confidence, consider adjusting parameters or target sequence
In the original benchmarking, pseudo-perplexity showed statistically significant negative correlation (p < 0.01) with AlphaFold-Multimer structural metrics (ipTM and pLDDT scores), validating its use as a selection criterion.
How does PepMLM work?
PepMLM adapts the ESM-2 protein language model for conditional peptide generation through a novel masking strategy. The approach treats binder design as a sequence reconstruction problem where peptides are predicted given their target protein context.
Masked language modeling
The core innovation positions peptide sequences at the C-terminus of target proteins during training. The entire binder region is masked, forcing the model to reconstruct it based solely on the target sequence context. This contrasts with traditional masked language models that mask random tokens throughout the sequence.
During inference, the model receives a target protein sequence concatenated with a masked peptide template of length . Top-k sampling with categorical probability distributions generates amino acids position-by-position, producing diverse binder candidates rather than a single deterministic prediction.
Training methodology
PepMLM fine-tunes ESM-2 (650M parameters) on merged datasets from PepNN and Propedia, comprising 10,000 training samples and 203 test sequences. The model constrains binders to maximum 50 amino acids and target proteins to 500 residues.
The training objective minimizes masked language modeling loss exclusively on binder regions:
where represents amino acids in the masked binder region, is the target protein sequence, and includes already-generated positions.
Benchmark performance
In comparative studies, PepMLM achieved superior performance against RFdiffusion on structured targets:
- Hit rate — >38% success rate compared to RFdiffusion's <30%
- Binding affinity — Top peptides demonstrated significantly stronger binding to NCAM1 than all RFdiffusion candidates
- Structural validation — AlphaFold-Multimer predictions confirmed binding pockets matching validated test sequences
Limitations
- Linear peptides only — Does not design cyclic peptides, stapled peptides, or structures with disulfide bridges
- No structural optimization — Generated sequences represent initial candidates that may benefit from cyclization, stapling, or other post-design modifications
- Sequence-only conditioning — Cannot incorporate structural constraints or binding site information when available
- Unknown amino acid tokens — Some positions may generate
X(any amino acid), requiring manual substitution or experimental screening - No affinity prediction — Pseudo-perplexity indicates sequence likelihood but does not directly predict binding affinity or values