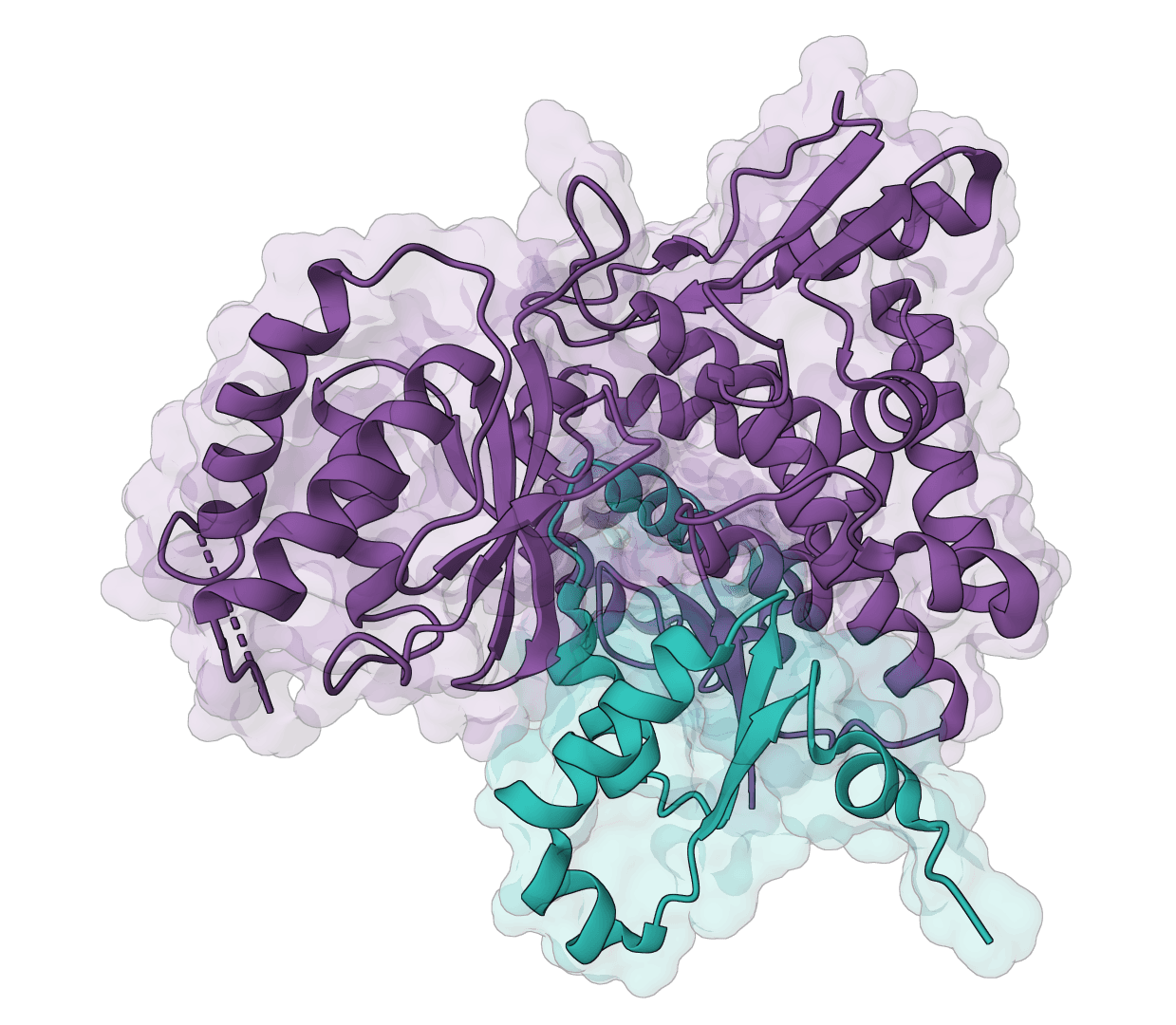What is IntelliFold 2?
IntelliFold 2 is an all-atom biomolecular structure prediction model for proteins, ligands, DNA, RNA, and mixed complexes. It extends the earlier IntFold foundation model with an updated architecture and a faster v2-Flash inference path, while retaining the project's focus on controllable prediction for practical design and screening workflows.
The IntFold technical report describes the platform as a controllable foundation model that reaches AlphaFold 3 level performance on broad biomolecular benchmarks while adding mechanisms for structural constraints, allosteric-state guidance, and downstream ranking. The IntelliFold 2 release positions the newer model family as an architectural refinement intended to improve accuracy on recent FoldBench evaluations.
Applications
- Protein structure prediction: Single-chain and multimer folding from sequence input.
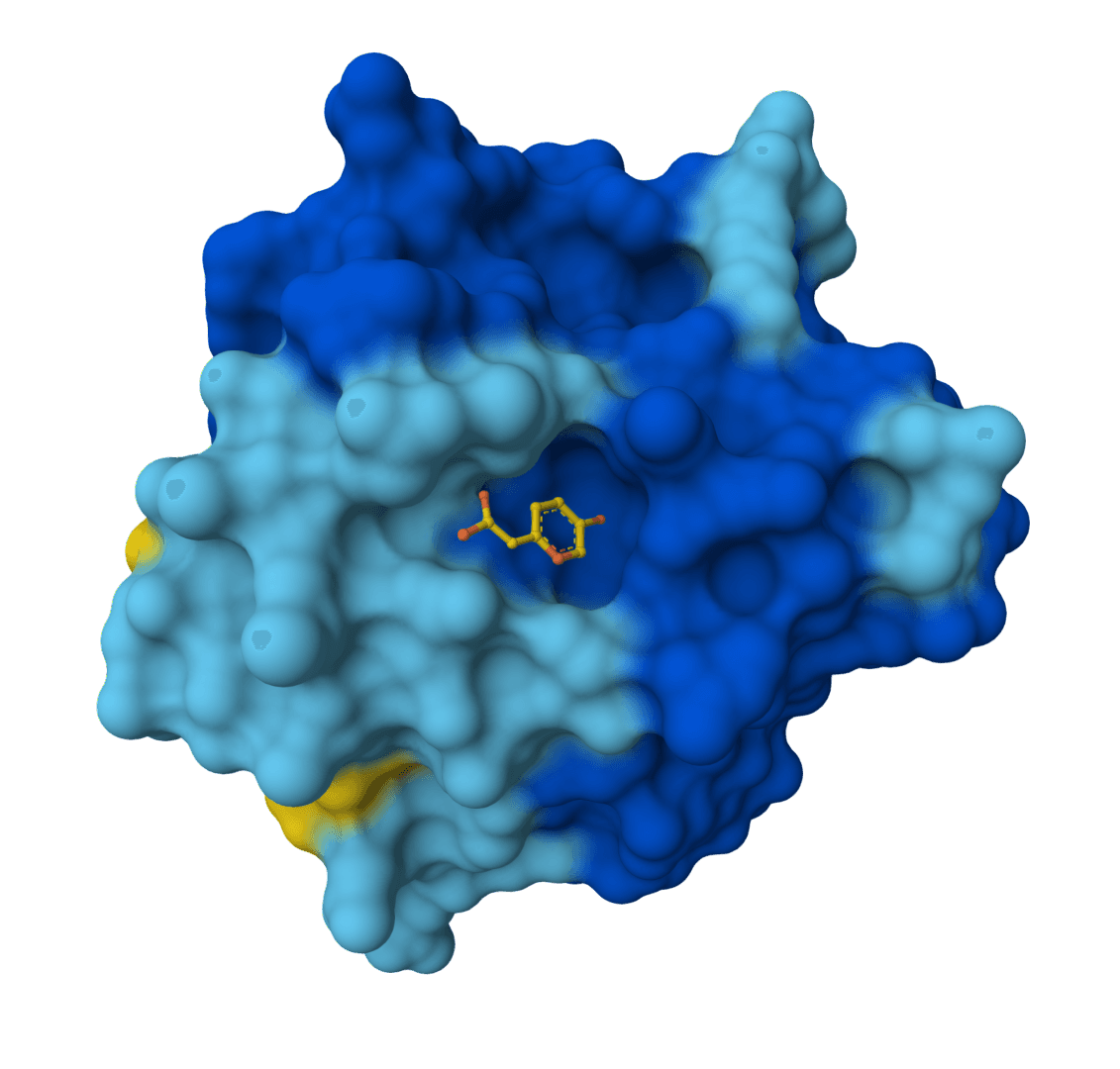
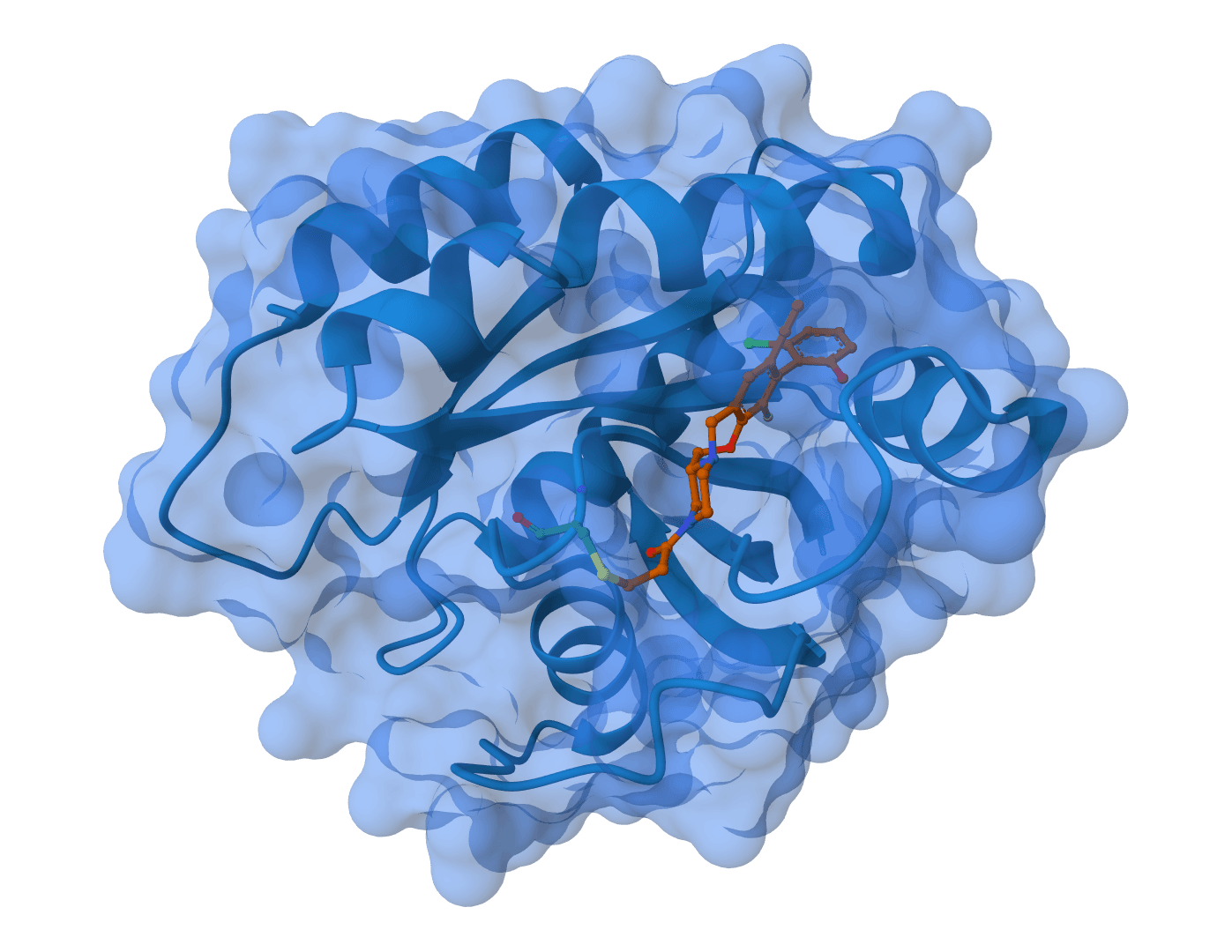
- Protein-ligand modeling: Joint prediction of receptor and ligand geometry in one run.
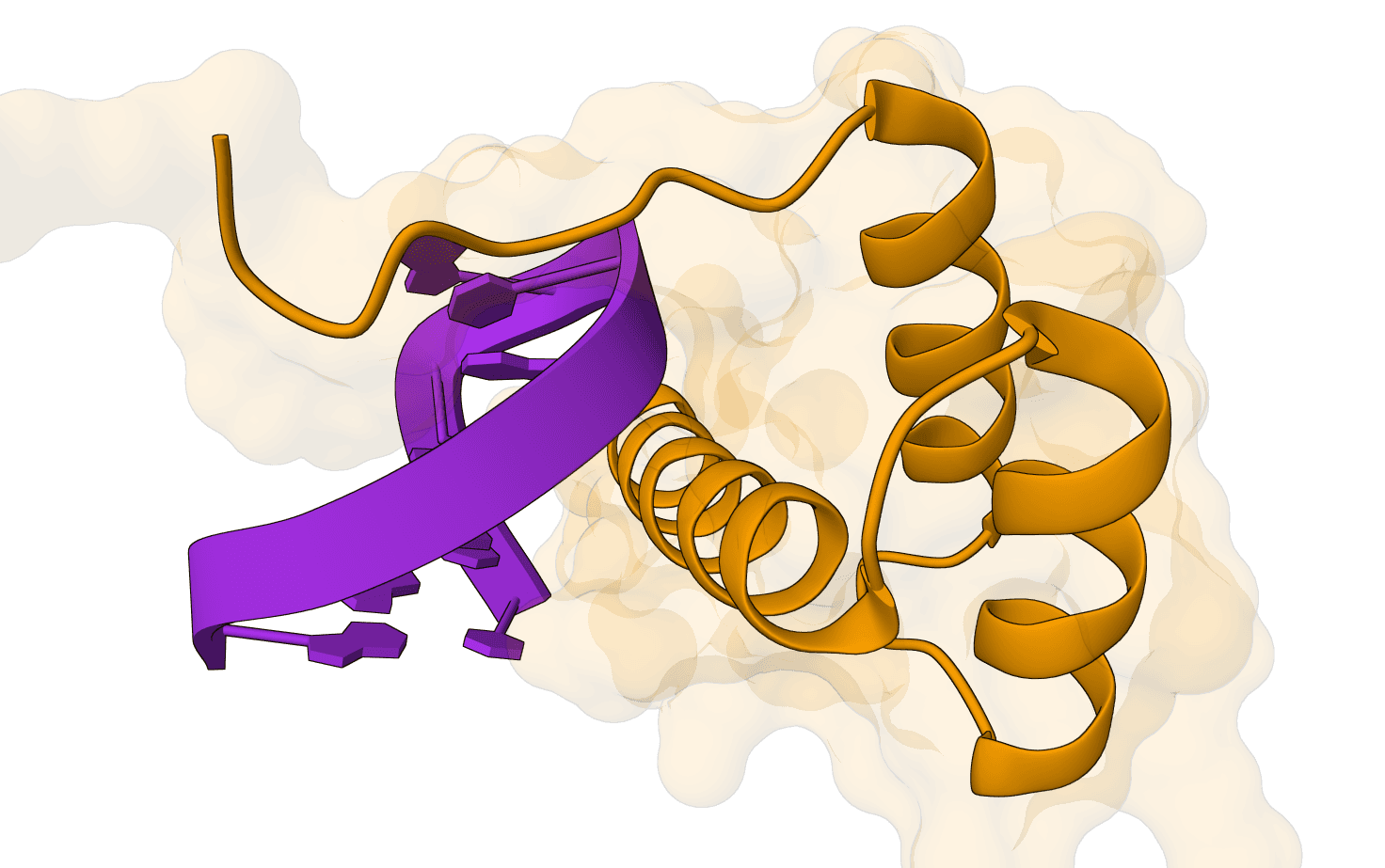
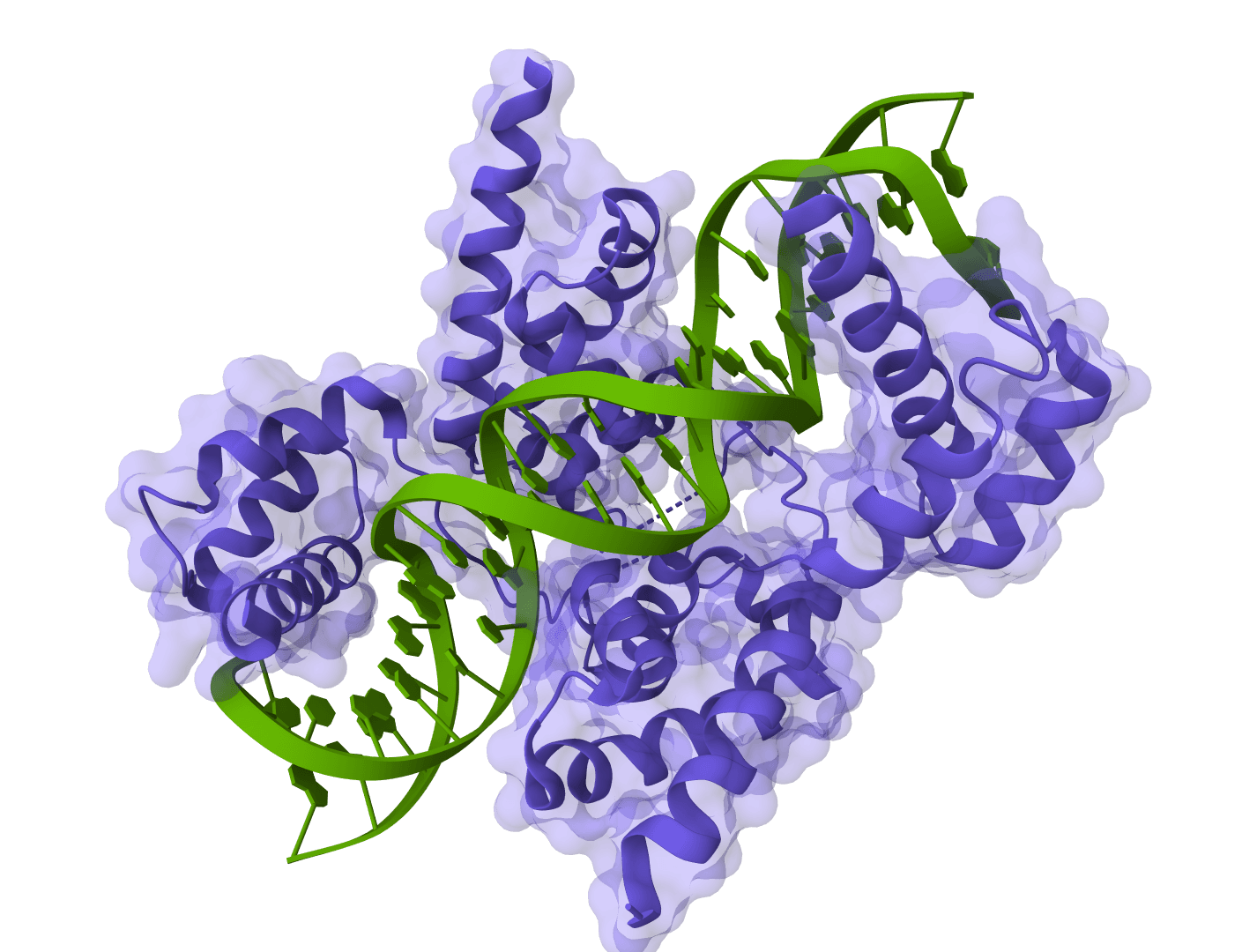
- Protein-nucleic acid complexes: Modeling systems that include DNA, RNA, or both.
- Conformation sampling: Generating multiple structures across seeds and diffusion samples to inspect alternative states.
How to use IntelliFold 2 online
ProteinIQ provides browser-based access to IntelliFold 2, so the model can be run without local GPU setup, Python installation, or manual environment configuration. Jobs are submitted from a single form that combines molecular inputs, MSA options, and inference settings.
At least one foldable biopolymer is required in practice: a protein, DNA, or RNA chain. The ProteinIQ deployment accepts up to 10 entries for each molecule type, with a combined maximum of 3,000 residues across all protein, DNA, and RNA chains.
Settings
Prediction parameters
MSA options
Output
ProteinIQ returns an interactive structure viewer, a downloadable results table, and the full file bundle for each run.
How does IntelliFold 2 work?
The IntFold project is built as an all-atom foundation model for biomolecular structure prediction. In the authors' description, its distinguishing feature is controllability: the same core system is designed to support general structure prediction as well as guided tasks such as constrained inference, alternative-state modeling, and binding-focused applications.
Controllable all-atom prediction
The IntFold technical report describes specialized adapters that can bias the model toward user-defined structural constraints, allosteric states, and affinity-related downstream tasks. That makes IntelliFold different from sequence-to-structure systems that only return an unconstrained best guess.
Diffusion sampling and recycling
ProteinIQ supports the model's diffusion and recycling controls directly. Diffusion sampling starts from noisy coordinates and refines them over many denoising steps, while recycling passes intermediate predictions back through the network for further correction. Increasing either parameter usually improves search depth, but the trade-off is longer runtime.
MSA-assisted or single-sequence inference
IntelliFold 2 can use server-generated MSAs through MMseqs2 or run from sequence input with MSA generation disabled. In protein systems with clear evolutionary homologs, MSA depth usually improves structural confidence; disabling MSA is more appropriate for speed-sensitive runs or synthetic targets with weak sequence family coverage.
Ensemble ranking
The IntFold paper also describes a training-free, similarity-based ranking strategy for selecting better predictions from an ensemble. On ProteinIQ, that ensemble is controlled indirectly through Diffusion samples and Seeds, which determine how many candidate structures are generated before ranking.
Interpreting results
IntelliFold 2 predictions are typically examined as a ranked set rather than a single definitive structure. Confidence metrics included in the IntelliFold 2 release materials and ProteinIQ configuration are the most useful first-pass filters.
For exploratory jobs, multiple seeds and diffusion samples are often more informative than a single large run with fixed randomness. If top-ranked structures differ substantially from one another, the target may be flexible, weakly constrained by sequence evidence, or genuinely multimodal.
Limitations
- IntelliFold 2 is still a predictive model rather than an experimental measurement, so flexible regions, disordered tails, and weakly supported interfaces should be interpreted cautiously.
- The open-source repository notes that the model natively supports templates, but the published inference pipeline adapted from Boltz-1 does not currently process templates. ProteinIQ therefore supports sequence, ligand, and MSA controls rather than template-guided prediction.
- Very large complexes remain constrained by sequence-length scaling and runtime. ProteinIQ caps total protein, DNA, and RNA residues at 3,000 for this deployment.
- MSA quality still matters for protein-heavy jobs. Predictions for de novo designs or sequences with few homologs may be less stable than predictions for well-covered families.