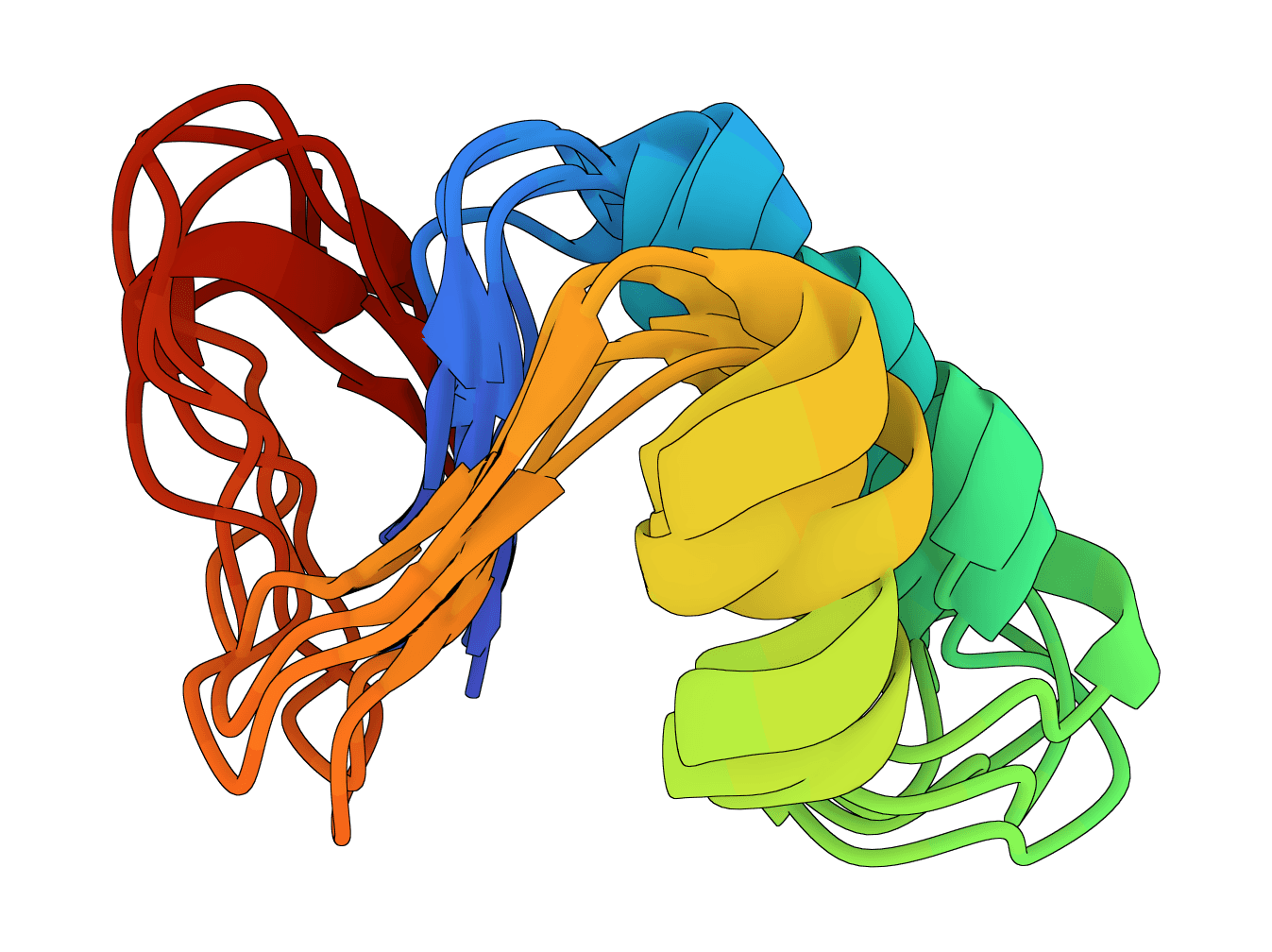Human Ubiquitin (76 aa) - Flexible regions
Related tools
AlphaFold2
AlphaFold2 via ColabFold for high-accuracy protein structure prediction. Uses MMSeqs2 API for MSA generation with no local databases required. Supports monomer and multimer prediction.
Boltz-2
Boltz-2 is a biomolecular foundation model for structure and binding affinity prediction. Supports proteins, ligands, DNA, and RNA in multi-component complexes. Automatically scales GPU resources for large complexes. Predicts binding affinity with near-FEP accuracy at 1000x faster speed.
Chai-1
Chai-1 is a multi-modal foundation model for molecular structure prediction. Predicts 3D structures for proteins, ligands, DNA, RNA, and multi-component complexes with high accuracy.
ESMfold
ESMfold is a fast, single-sequence protein structure predictor from Meta AI. Predicts 3D protein structures directly from amino acid sequences without requiring multiple sequence alignments (MSA), making it significantly faster than AlphaFold while automatically scaling GPU resources for larger proteins.
ImmuneBuilder
ImmuneBuilder predicts 3D structures of immune receptor proteins including antibodies, nanobodies, and T-cell receptors. It uses ABodyBuilder2, NanoBodyBuilder2, and TCRBuilder2/TCRBuilder2+ to generate structures with per-residue error estimates and optional ensemble artifacts.
IntelliFold 2
Controllable biomolecular structure prediction model for proteins, ligands, DNA, RNA, and multi-component complexes. IntelliFold 2 supports fast v2-Flash inference, optional MSA generation, and ranked confidence outputs.
LMI4Boltz
LMI4Boltz is a low-memory fork of Boltz for biomolecular structure and binding affinity prediction. It preserves Boltz inference behavior while reducing VRAM use with in-place pair updates, CPU offload, reduced precision pair representation, and aggressive chunking.
MiniFold
MiniFold is a fast single-sequence protein structure predictor that is 10-20x faster than ESMFold. It predicts 3D protein structures directly from amino acid sequences without requiring multiple sequence alignments (MSA), making it ideal for rapid structure prediction.
OpenFold-3
OpenFold-3 is an open-source AI model for biomolecular structure prediction, aiming to reproduce AlphaFold3. Predicts 3D structures for proteins, RNA, DNA, and small molecule ligands with high accuracy.
Protenix
Open-source AlphaFold 3 implementation by ByteDance for biomolecular structure prediction. Predicts 3D structures for proteins, RNA, DNA, and small molecule ligands with high accuracy.
What is AlphaFlow?
Protein structures in the PDB and in molecular dynamics simulations rarely describe a single rigid shape. Loops open and close, termini wander, domains breathe, and binding sites can appear only in a subset of conformations. AlphaFlow models that variability by turning AlphaFold-style structure prediction into a generative ensemble method.
AlphaFlow, developed by Jing, Berger, and Jaakkola, includes two model families. AlphaFlow uses AlphaFold with multiple sequence alignments. ESMFlow uses ESMFold and runs from sequence alone. ProteinIQ supports both model families: ESMFlow for single-sequence ensemble generation, and AlphaFlow for MSA-backed AlphaFold-style ensemble generation.
For a single best-guess structure, ESMFold or AlphaFold 2 is usually the clearer choice. AlphaFlow is useful when the question is about plausible conformational spread rather than one static model.
How to use AlphaFlow online
AlphaFlow runs online in ProteinIQ by selecting an input mode, providing one protein sequence, choosing a matching model variant, and generating a conformational ensemble as downloadable PDB files. ESMFlow mode needs only a sequence. AlphaFlow mode accepts the same sequence plus an optional .a3m multiple sequence alignment, and generates the alignment automatically when none is uploaded.
Inputs
| Input | Accepted format | Notes |
|---|---|---|
Protein sequence | FASTA, raw amino acid sequence, .fasta, .fa, or .txt | One protein sequence between 10 and 1600 residues. Standard amino acids are expected. |
Pre-computed MSA | Optional .a3m | Appears only after AlphaFlow input mode is set to AlphaFlow. The uploaded alignment must be valid A3M with the query sequence first. Invalid .a3m files fail clearly rather than falling back to generated MSA. |
RCSB fetch | PDB ID, for example 1UBQ | ProteinIQ fetches FASTA from RCSB when a PDB ID is used. The resulting sequence, not the PDB coordinates, is used for ESMFlow inference. |
Job name | Text | Optional label for finding the run later. |
AlphaFlow on ProteinIQ is designed for single-chain protein sequences. Multimer interfaces, ligand-bound states, membranes, cofactors, and chain-specific contacts are not modeled directly from the input. When a PDB ID is fetched from RCSB, the sequence is used as the input and the deposited coordinates are not used as a structural template.
A3M validation
Uploaded .a3m alignments are checked before the job starts. ProteinIQ accepts permissive A3M content, but rejects files that would make the AlphaFold-style feature pipeline ambiguous.
| Check | Requirement |
|---|---|
| FASTA headers | Each record starts with a non-empty header line beginning with >. |
| Sequence records | Every header has at least one sequence line. |
| Characters | Sequences use plausible A3M characters: amino acid letters, lowercase insertion letters, -, ., or *. |
| Aligned length | After stripping lowercase insertion characters, every record has the same aligned column length. |
| Query sequence | The first record has a non-empty ungapped query sequence after removing lowercase insertions and gap-like characters. |
If no .a3m is uploaded in AlphaFlow mode, ProteinIQ generates an MSA server-side with an MMseqs2 workflow compatible with ColabFold-style A3M output. The generated alignment is returned with the results so the run can be inspected or repeated.
Before running
- Trim expression tags and purification handles: Flexible His-tags or long artificial linkers can dominate the ensemble and make biologically relevant motion harder to see.
- Keep domains interpretable: Multi-domain proteins can be useful, but long flexible linkers may cause large global rearrangements. Running individual domains can make local flexibility easier to inspect.
- Check the sequence alphabet: Ambiguous residues such as
X,B,Z, orUshould be resolved before inference when possible. - Start with 10 samples: The default gives enough structures to spot repeated motion without making the first run slow. Increase to 25 or 50 when comparing domain movement, loop states, or binding-site exposure.
Human ubiquitin (1UBQ, 76 residues) is a useful sanity-check target because the beta-grasp fold should remain compact while termini and surface loops vary modestly across samples. A run where the core unfolds or most samples look unrelated is a warning sign to inspect the input sequence and settings.
Model settings
| Setting | Default | Description |
|---|---|---|
AlphaFlow input mode | ESMFlow | Controls both the input form and the available model variants. ESMFlow uses sequence only. AlphaFlow shows the optional .a3m upload and otherwise generates an MSA server-side. |
Model variant | ESMFlow MD Base in ESMFlow mode, AlphaFlow MD Base in AlphaFlow mode | Selects the checkpoint. The menu is scoped to the selected input mode, so ESMFlow variants appear only in ESMFlow mode and AlphaFlow variants appear only in AlphaFlow mode. |
Number of samples | 10 | Number of PDB conformations to generate, from 1 to 50. More samples give a better view of ensemble spread, but runtime scales roughly linearly with sample count. |
MD models learn conformational variation from molecular dynamics trajectories. PDB models learn diversity across experimental structures. Base models are the full checkpoints. Distilled models are faster and automatically use the distilled inference settings.
Choosing a model variant
| Variant | Best fit | Practical tradeoff |
|---|---|---|
ESMFlow MD Base | Studying thermal flexibility, mobile loops, termini, and domain breathing around physiological conditions | Recommended default. Slower than distilled models, but closest to the full MD-trained ESMFlow behavior. |
ESMFlow PDB Base | Looking for experimental-state diversity, such as alternative crystal forms or condition-dependent conformations | Can reflect broader structural heterogeneity than MD-like local motion. |
ESMFlow MD Distilled | Fast exploratory ensemble generation when many sequences need screening | Faster, with some accuracy loss relative to the base model. Uses No diffusion and Noisy first automatically. |
ESMFlow PDB Distilled | Fast comparison of experimental-ensemble-like conformations | Useful for triage, not the best choice for final structural interpretation. |
AlphaFlow MD Base | MSA-backed molecular-dynamics-like ensembles | Uses AlphaFold-style MSA features. Slower and more expensive than ESMFlow, but closer to the original AlphaFlow method. |
AlphaFlow MD Distilled | Faster MSA-backed MD-like exploratory ensembles | Uses the distilled inference mode. |
AlphaFlow PDB Base | MSA-backed experimental-state ensemble diversity | Enables Self conditioning and MSA resampling by default for PDB-style sampling. |
AlphaFlow PDB Distilled | Faster MSA-backed PDB-like ensembles | Uses PDB-style sampling plus the distilled inference mode. |
Advanced settings
| Setting | Default | Description |
|---|---|---|
Inference steps | 10 | Number of denoising steps, from 2 to 50. The standard setting is 10. Lower values truncate generation and can reduce diversity. Higher values cost more runtime. |
Diversity (tmax) | 1.0 | Starting point in the generation schedule, from 0.2 to 1.0. 1.0 samples from the full schedule. Lower values keep samples closer together. |
Self conditioning | Off | Enables --self_cond. AlphaFlow PDB variants enable it automatically for PDB-style sampling. |
No diffusion | Off | Enables --no_diffusion. Distilled variants enable this automatically. |
Noisy first | Off | Enables --noisy_first. Distilled variants enable this automatically. |
The two controls that most users should change first are Number of samples and Model variant. Inference steps, Diversity (tmax), and the boolean inference flags are better treated as reproducibility controls. Changing them changes the sampling procedure, not just the display.
Results
ESMFlow returns one primary PDB file per sample. AlphaFlow returns a canonical multi-model PDB containing the full sampled ensemble, plus individual sample PDB files for convenient viewing. When AlphaFlow uses an uploaded or generated .a3m, that alignment is returned as a secondary downloadable file for reproducibility.
| Output | Meaning |
|---|---|
sample_01.pdb, sample_02.pdb, ... | Individual predicted conformations in PDB format. Each file is a complete protein model for the same input sequence. |
{name}.pdb | AlphaFlow modes only. Multi-model PDB containing the sampled ensemble. |
{name}.a3m | AlphaFlow modes only. Uploaded or generated MSA used for inference. |
Model variant | Checkpoint used for the run, such as esmflow_md_base or esmflow_pdb_distilled. |
Sample index | Position of the sampled conformation in the generated ensemble. It is not a confidence rank. |
Inference settings | steps, tmax, self_cond, no_diffusion, and noisy_first are recorded so runs can be compared later. |
AlphaFlow does not return a pLDDT confidence table or a binding score. The value of the run is in the ensemble itself: how similar the structures are, which regions move, and whether the sampled conformations suggest more than one plausible state.
| Pattern in the ensemble | Likely interpretation | Follow-up check |
|---|---|---|
| Core secondary structure overlaps across most samples | The model has a consistent fold hypothesis | Inspect loops, termini, and domain interfaces rather than whole-protein RMSD alone. |
| One loop repeatedly opens or closes | The loop may be mobile or weakly constrained by sequence context | Compare with known active-site loops, disorder predictions, or mutagenesis data. |
| Two domains stay folded but change relative orientation | Hinge-like motion may be plausible | Align one domain first, then measure displacement of the other domain. |
| Only one sample shows an extreme rearrangement | The structure is more likely an outlier than a stable alternate state | Look for the same movement in additional samples before using it for docking or design. |
| Most samples are globally inconsistent | The input may be too long, disordered, or outside the model's reliable regime | Try domain-level constructs or compare a single-structure prediction with ESMFold. |
Interpreting AlphaFlow ensembles
An AlphaFlow output should be read as a set of plausible conformations, not as a ranked list where sample 1 is best. The sample index only records generation order.
Large differences across samples usually point to regions where the model sees conformational uncertainty or mobility. In practice, the most useful checks are visual:
- Stable core: Helices and sheets that overlap across samples are more likely to represent a consistent fold.
- Flexible loops and termini: Disordered ends and solvent-exposed loops often spread widely. That spread is expected and should not be treated as a modeling failure by itself.
- Domain motion: If two domains remain internally stable but shift relative to each other, the ensemble may suggest hinge-like motion.
- Binding-site exposure: A pocket that appears in only some samples can be useful for hypothesis generation, especially before docking or mutagenesis planning.
- Outlier samples: A single highly distorted sample should not outweigh the rest of the ensemble. Inspect whether the same movement appears repeatedly.
Quantitative RMSD analysis is useful after the job, but RMSD needs a reference choice. Whole-protein RMSD can be dominated by flexible tails. For many proteins, aligning the stable core first and then measuring loop or domain movement gives a clearer interpretation.
How AlphaFlow works
AlphaFlow trains structure predictors as flow-matching models. During training, the model sees an interpolation between a noisy protein backbone representation and a target structure:
Here, is sampled from a harmonic prior, is the target structure, and marks progress along the denoising path. At inference time, the model starts from a noisy configuration and repeatedly predicts a cleaner structure along the schedule.
ESMFlow applies the same idea to ESMFold. Because ESMFold is a single-sequence model, ESMFlow can run without MSA generation. That makes it practical for fast web execution and for proteins where homologous sequence coverage is weak.
AlphaFlow mode uses AlphaFold-style MSA features. A supplied .a3m file is used directly after validation. Without an upload, ProteinIQ generates the alignment first, then writes it into the directory layout expected by the AlphaFlow inference code before sampling structures. That extra MSA step is the main reason AlphaFlow mode is slower than ESMFlow mode.
Training data behind the model variants
| Training set | What it captures | Interpretation consequence |
|---|---|---|
MD | Molecular dynamics trajectories from ATLAS at 300K | Better for local thermal motion, loop mobility, and dynamic fluctuations around an existing fold. |
PDB | Experimental structures deposited under different conditions | Better for alternative crystallographic or cryo-EM states, ligand-bound differences, and broader experimental heterogeneity. |
The distinction matters. An MD-trained model may emphasize physically local fluctuations. A PDB-trained model may sample larger state changes if similar state diversity appears in experimental structures.
When to use AlphaFlow vs alternatives
| Tool | Best use | Difference from AlphaFlow |
|---|---|---|
| AlphaFlow | Generating a conformational ensemble from one protein sequence | Produces multiple plausible structures rather than one best structure. |
| ESMFold | Fast single-structure prediction without MSA generation | Better when a single model and pLDDT-style confidence are needed. |
| AlphaFold 2 | High-accuracy structure prediction where MSA information is helpful | Usually stronger for a single final structure, but not designed to sample an ensemble. |
| AF-Cluster | Exploring alternate AlphaFold conformations by clustering subsampled MSA predictions | Useful when MSA diversity may reveal different states, but it still depends on AlphaFold-style prediction runs. |
| Boltz-2 | Structure prediction for complexes involving proteins, ligands, DNA, or RNA | Better for multimolecular complexes and binding affinity workflows. |
| MD trajectory analysis | Measuring flexibility from an existing molecular dynamics trajectory | Works from simulated trajectory data instead of generating conformations from sequence. |
AlphaFlow fits best between structure prediction and simulation. It can suggest flexible regions worth checking before setting up molecular dynamics, docking, mutagenesis, or construct design. It should not replace experimental dynamics data when NMR, HDX-MS, cryo-EM heterogeneity, or high-quality MD trajectories are available.
Practical limits
- MSA generation can dominate runtime: AlphaFlow modes require MSA features. Uploaded
.a3mfiles make runs more reproducible; automatic MMseqs2 generation depends on external search availability. - No ligand or multimer conditioning: The input is a protein sequence. Ligands, cofactors, membranes, partner chains, and DNA or RNA binding partners are not included in generation.
- Backbone-centered ensemble interpretation: The method is most useful for backbone conformational diversity. Side-chain packing and small binding-site rearrangements need follow-up validation.
- Long sequences cost more: Memory and runtime increase sharply with sequence length. ProteinIQ accepts up to 1600 residues, but shorter constructs are usually easier to interpret.
- Samples are hypotheses: Repeated motion across several samples is more informative than a single unusual conformation. Experimental or simulation evidence is still needed before claiming a functional state.