Input
Your jobs will appear here. To create a job, sign up below.
Sign up
MolProbity
Validate protein structure quality with all-atom contact analysis, Ramachandran plots, rotamer assessment, and geometry checks.
Ligand fixer
Fix ligand files that fail RDKit, Meeko, or docking preparation. Repair SDF, MOL, and MOL2 inputs, apply safe chemistry cleanup, and export docking-ready SDF files.
PDB2PQR
PDB2PQR prepares protein structures for electrostatics calculations by adding missing atoms, predicting protonation states using PROPKA, and assigning atomic charges and radii from standard force fields.
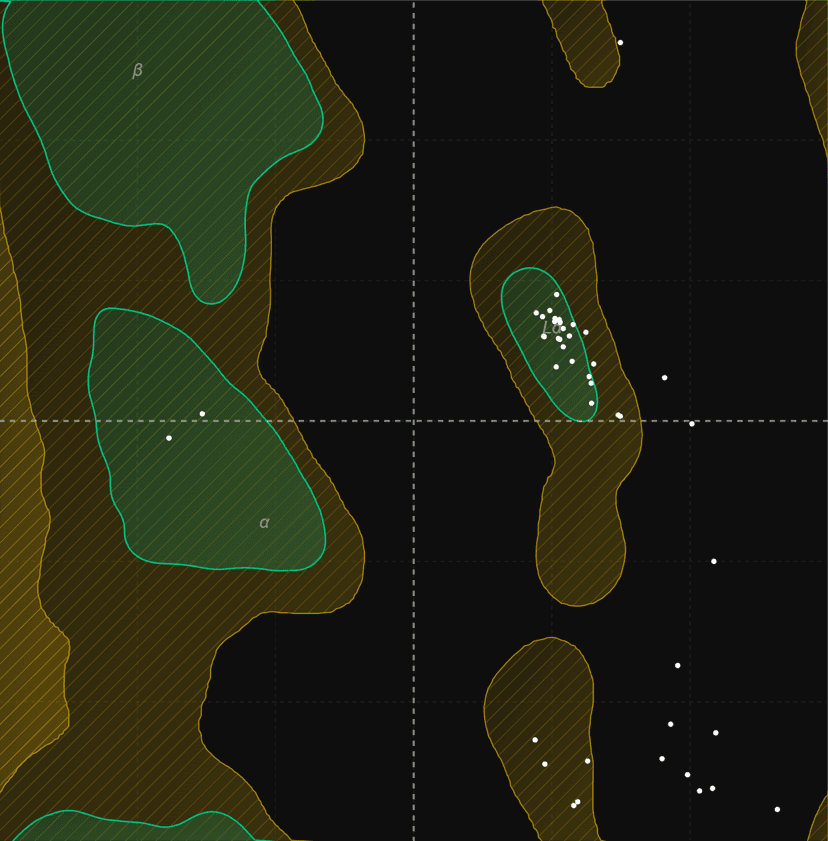
Ramachandran plot
Generate Ramachandran plots from PDB structures to analyze protein backbone dihedral angles (phi/psi). Visualize favored, allowed, and outlier regions.
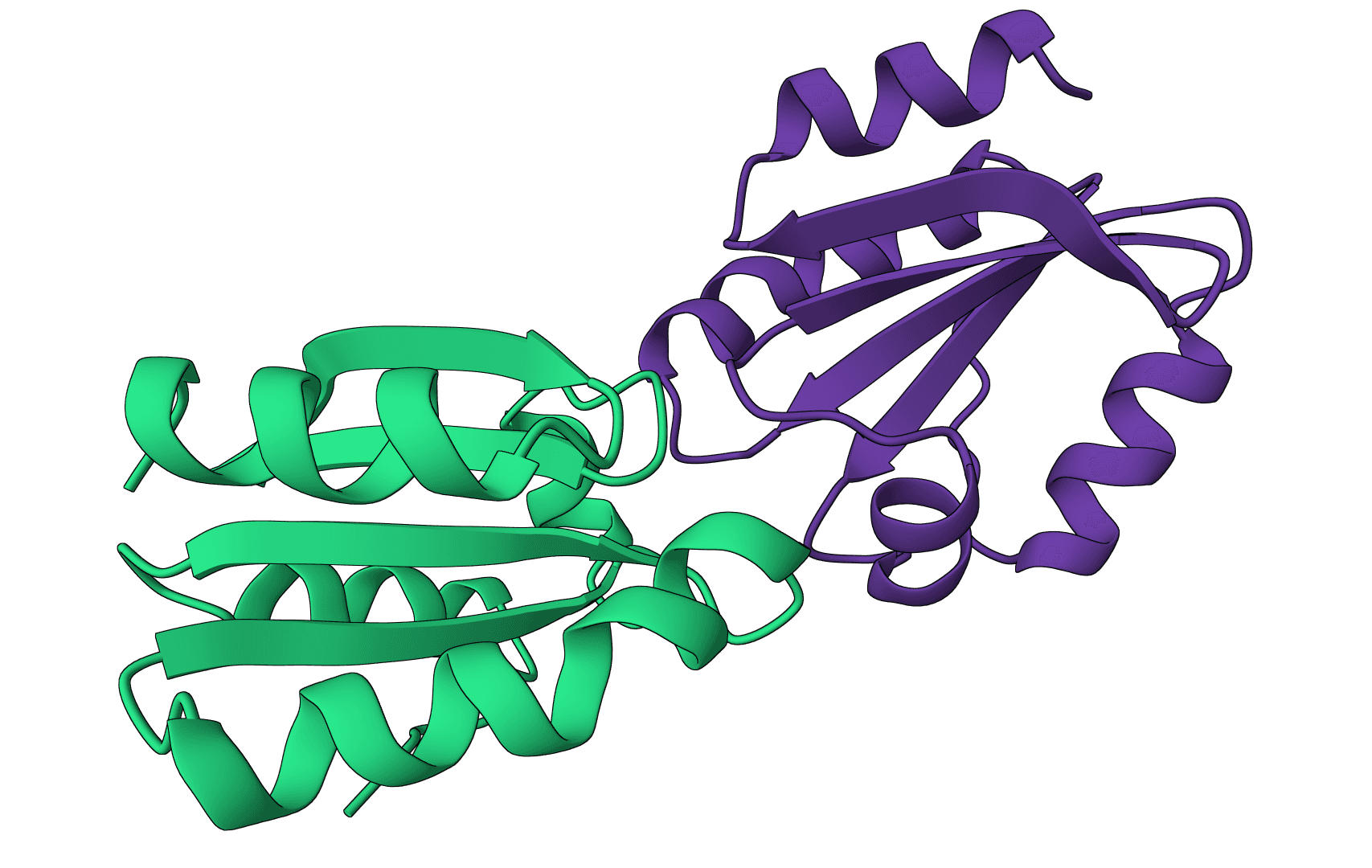
DockQ
Assess docking model quality by comparing predicted complexes against native references. DockQ v2.1.3 supports protein, nucleic-acid, and supported small-molecule interfaces with faithful native metrics.
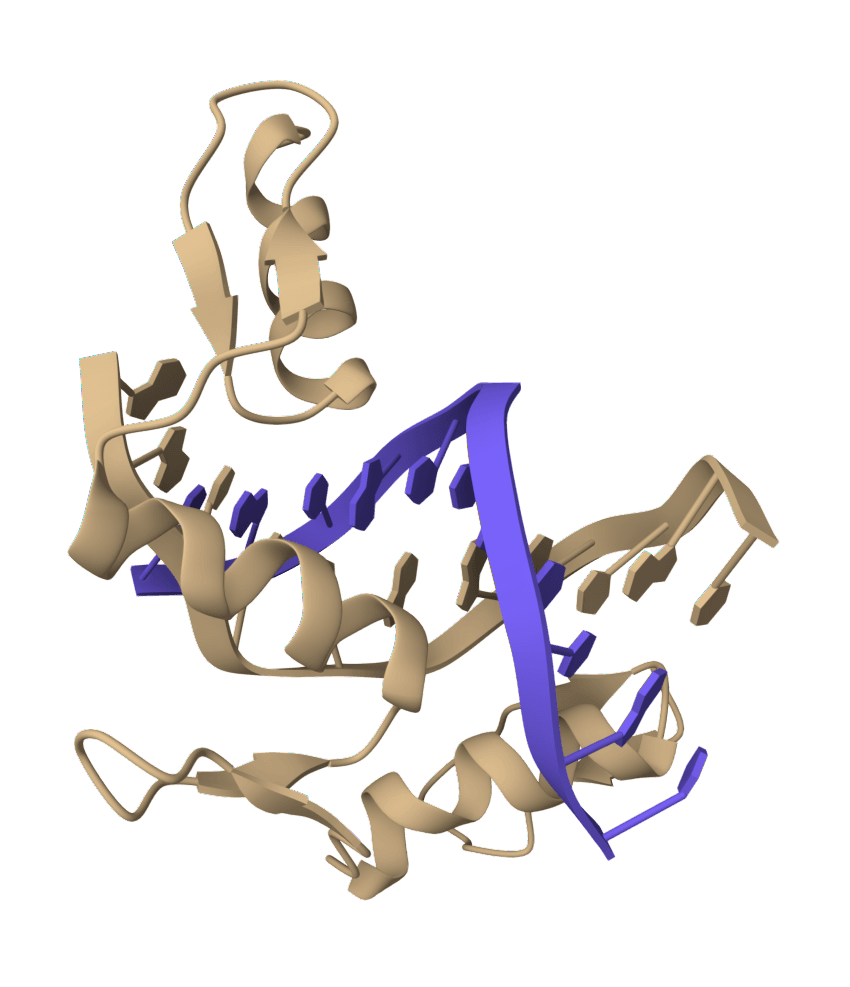
Filter DNA
Clean and filter DNA sequences by removing or replacing non-standard nucleotide characters. Supports multiple filter modes including standard 4 bases, IUPAC ambiguity codes, and custom character sets.
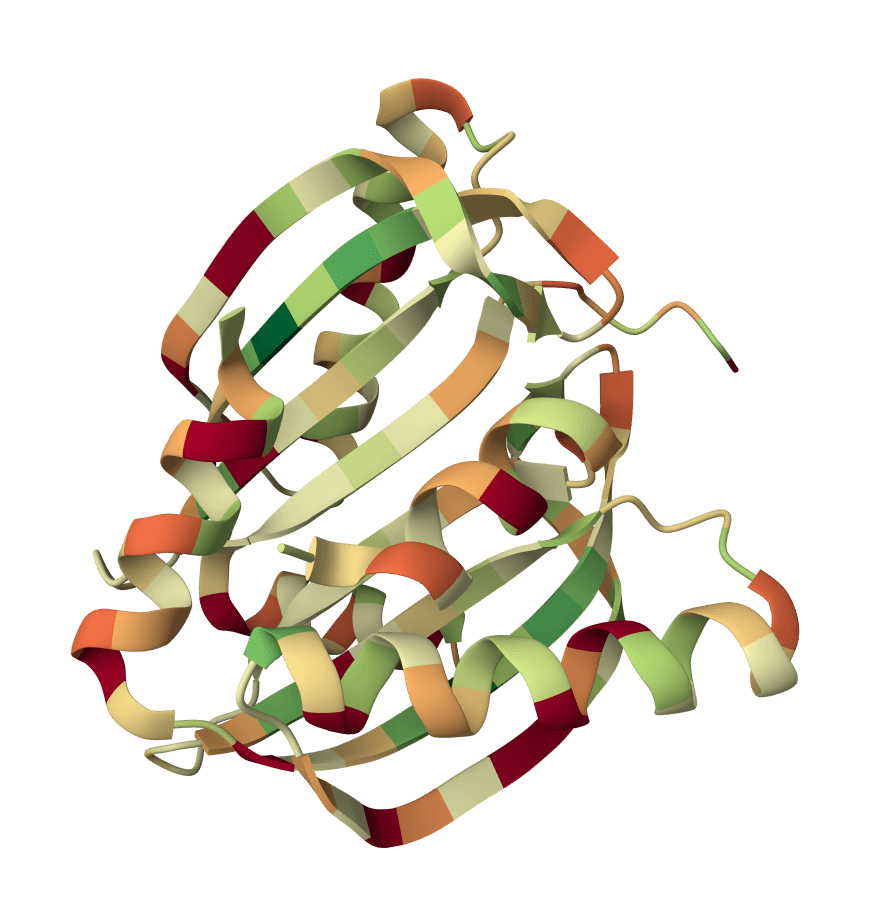
IPSAE
Scoring function for interprotein interactions in AlphaFold2, AlphaFold3 and Boltz predictions. Calculates ipSAE, ipTM, pDockQ, pDockQ2, and LIS scores to assess protein-protein interface quality.
PDB to CIF Converter
Convert Protein Data Bank files to Crystallographic Information File format

PDB to FASTA converter
Convert Protein Data Bank files to FASTA sequence format
PDB to MOL2 Converter
Convert Protein Data Bank files to MOL2 molecular format
What is PDB Fixer?
PDB Fixer is an open-source tool developed as part of the OpenMM molecular simulation toolkit. It automatically repairs Protein Data Bank files in both PDB and mmCIF/PDBx formats, fixing structural problems that prevent successful molecular dynamics simulations.
ProteinIQ's web implementation can process up to 10 uploaded or fetched structures in one job with one shared settings block, then return one fixed PDB per input together with a summary.
Experimental protein structures from X-ray crystallography or cryo-EM frequently contain issues: hydrogen atoms aren't resolved, side chains or entire loops may be missing due to flexibility, modified residues need conversion, and crystallization artifacts (buffers, salts, duplicate chains) clutter the file. These problems cause simulation software like GROMACS, Amber, or OpenMM to fail.
PDB Fixer addresses eight primary structural problems:
- Missing hydrogens: Adds all hydrogen atoms absent from X-ray structures
- Missing heavy atoms: Completes side chain atoms in regions with poor electron density
- Missing terminal atoms: Adds capping atoms at chain ends
- Missing residues: Builds complete loops in disordered regions
- Nonstandard residues: Converts modified amino acids (e.g., selenomethionine) to standard equivalents
- Heterogens: Removes or keeps ligands, ions, and water molecules
- Alternate locations: Selects single conformations when multiple exist
- Solvent/membrane: Constructs water boxes or lipid bilayers for explicit solvent simulations
After fixing your structure, use PDB Viewer to visually inspect the results or PDB to FASTA to extract sequences.
How does PDB Fixer work?
PDB Fixer operates as a stepwise pipeline that analyzes molecular topology and atomic coordinates. It uses residue templates from the PDB Chemical Component Dictionary to identify what atoms should exist and add any that are missing.
Template-based correction
For each residue, PDB Fixer compares the atoms present against the expected template. Missing atoms are placed using ideal bond lengths, angles, and dihedral values from the template library. This ensures chemically reasonable starting positions.
Energy minimization
After adding atoms, the tool runs brief energy minimization using OpenMM's force fields. This resolves steric clashes between newly added atoms and the existing structure while preserving experimentally determined coordinates as much as possible.
Protonation states
When adding hydrogens, PDB Fixer assigns protonation states based on the specified pH. Titratable residues (histidine, aspartate, glutamate, lysine) adopt appropriate charge states. Special cases like disulfide bonds and histidine tautomers are handled automatically.
GPU acceleration for large structures
Structures exceeding 10,000 atoms are automatically routed to GPU-accelerated processing (NVIDIA T4 with CUDA-enabled OpenMM). Energy minimization during atom placement is the main computational bottleneck, and GPU offloading reduces processing time significantly for large complexes. Smaller structures are processed on CPU, where overhead is negligible and startup is faster. The routing is transparent — no configuration is needed.
Loop modeling
Missing residues are reconstructed using fragment-based loop building followed by minimization. The algorithm uses SEQRES records in the PDB file to identify which residues should exist, then builds them with idealized geometry before refining with energy minimization.
How to fix a PDB or mmCIF file using PDB Fixer online
Step 1: Load your structure
Upload one or more structure files or enter one or more RCSB PDB IDs (e.g., 1HSG for HIV-1 protease). Both PDB (.pdb, .ent) and mmCIF/PDBx (.cif, .mmcif, .pdbx) formats are accepted, up to 50 MB per file. The current ProteinIQ interface accepts up to 10 structures per job and applies the same repair settings to every entry in the run.
Step 2: Configure core processing options
Review the default settings under "Core processing." For most simulation preparation, keep heterogens set to Keep all to preserve ligands and ions. If you need to remove specific chains from a multi-chain complex, list their IDs in "Remove chains" (e.g., B, C).
Step 3: Select what to add
Under "Add options," the defaults enable "Add missing heavy atoms" and "Add missing hydrogens." These are essential for MD simulations. Enable "Add missing residues" only if you need complete loops—this significantly increases processing time for structures with large gaps.
Step 4: Add solvent (optional)
For explicit solvent MD simulations, enable "Add solvent box." Configure the ionic strength (typically 0.15 M for physiological conditions) and select your preferred box geometry. Dodecahedral boxes reduce computational cost compared to cubic boxes.
Step 5: Run and download
Click Fix Structure to start processing. Once complete, download the fixed PDB file directly or view it using the integrated PDB Viewer.
Multi-structure processing
Each input structure is processed separately, and the output list preserves the original input order.
When names are available, ProteinIQ keeps the original file names or RCSB identifiers for each repaired structure. If the incoming label is only a placeholder such as PDB 1, the platform attempts to recover a more useful name from the uploaded file name, the fetched identifier, or the PDB/mmCIF content itself.
PDB Fixer vs manual preparation
Many researchers prepare structures manually using tools like PyMOL, Chimera, or Swiss-PdbViewer. PDB Fixer offers several advantages for routine preparation tasks.
| Feature | PDB Fixer | Manual preparation |
|---|---|---|
| Input formats | PDB and mmCIF/PDBx | Varies by software |
| Missing atoms | Automatic detection and placement | Requires scripting or plugin |
| Hydrogens | pH-dependent protonation states | Often uniform or default states |
| Loop building | Automated with minimization | Requires homology modeling tools |
| Large structures | GPU-accelerated (>10k atoms) | CPU-only in most tools |
| Reproducibility | Deterministic with fixed seed | Depends on operator |
| Processing time | Minutes (GPU for large structures) | Hours for complex cases |
| Learning curve | Minimal | Requires software expertise |
PDB Fixer is ideal for routine multi-structure processing and simulation setup. For complex cases requiring manual intervention (e.g., unusual ligands, specific protonation states, or membrane positioning), use PDB Fixer as a starting point and refine manually.
Inputs & settings
Input requirements
Upload a PDB file (.pdb, .ent) or mmCIF/PDBx file (.cif, .mmcif, .pdbx), or fetch directly from RCSB using a PDB ID. Maximum file size is 50 MB per file, and up to 10 structures can be included in one job. mmCIF content is detected automatically by looking for data_ headers or _atom_site. tokens.
| Input mode | Description |
|---|---|
Upload | Accepts up to 10 structure files in PDB or mmCIF/PDBx format under one input card. |
RCSB fetch | Accepts multiple PDB identifiers in one submission and repairs each entry separately. |
Shared settings | All structures in the same job use one shared settings configuration. |
Core processing
- pH value: Sets protonation states for titratable groups when adding hydrogens. This setting only applies when "Add missing hydrogens" is enabled.
- Heterogens: Controls handling of non-protein molecules.
Keep allpreserves ligands, ions, and water from the original structure.Keep only waterremoves ligands and ions but preserves crystallographic waters.Remove allstrips everything except the protein chains. - Remove chains: Comma-separated chain IDs to remove (e.g.,
B, C, D). Useful for extracting a single chain from a multi-chain complex. - Apply mutations: Introduce point mutations during processing. Format:
CHAIN:ORIGINAL-POSITION-NEW(e.g.,A:ALA-57-GLY, B:VAL-123-ILE). This lets you study mutant variants without manual editing.
Add options
These settings control what structural elements PDB Fixer adds to your structure.
- Add missing residues: Builds entire missing loops from SEQRES records. Enable this when you need complete chains for simulation. Slow for structures with large gaps (10+ residue loops).
- Add missing heavy atoms: Completes truncated side chains. We recommend keeping this enabled for simulation preparation.
- Add missing hydrogens: Adds hydrogen atoms at the specified pH. Required for most MD simulations since X-ray structures lack hydrogens.
- Replace nonstandard residues: Converts modified amino acids (selenomethionine, phosphoserine, etc.) to their standard equivalents. Enable for standard force field compatibility; disable to preserve post-translational modifications.
Solvent box options
Adding explicit water creates a simulation-ready system. The protein is surrounded by a rectangular or truncated box of water molecules with counterions for charge neutralization.
- Add solvent box: Enables water box construction. Significantly increases processing time and output file size.
- Cation/Anion: Ion types for charge neutralization.
Na+/Cl-is standard for most simulations. ChooseK+for systems where potassium is physiologically relevant. - Ionic strength: Molar concentration of added salt.
0.15M matches physiological conditions. Increase for high-salt studies (halophiles, aggregation). - Box geometry: Shape of the periodic boundary box.
Cubicis standard and compatible with all software.DodecahedronandOctahedronreduce water count by ~30% while maintaining minimum distance from protein to box edge—faster simulations with equivalent accuracy. - Box sizing: Choose between automatic padding (distance from protein to box edges) or explicit X/Y/Z dimensions in nanometers.
- Water model: Select the water model for solvent.
TIP3Pis the standard choice for most force fields.TIP4P-Ewprovides improved density and diffusion properties.SPC/Eis popular for GROMACS workflows.
Membrane options
Membrane systems embed the protein in a lipid bilayer for studying membrane proteins (GPCRs, ion channels, transporters).
- Add lipid membrane: Constructs a membrane system. Cannot be combined with solvent box—the membrane system includes water and ions automatically.
- Lipid type: Composition of the bilayer.
POPC(palmitoyl-oleoyl-phosphatidylcholine) is the most common choice for general membrane protein studies.POPEis preferred for bacterial membranes. - Membrane center Z: Position of the bilayer center along the Z-axis in nanometers. Set to
0.0for automatic centering. Adjust when you need the protein positioned asymmetrically in the membrane. - Minimum padding: Distance from the protein to the box edges in nanometers.
Advanced options
- Force field: Select the molecular mechanics force field for atom placement and minimization.
AMBER14is recommended for most use cases.CHARMM36is preferred if you plan to run simulations with CHARMM-compatible software. - Random seed: Set a specific value for reproducible structure generation. Use
0for random initialization. Fixed seeds ensure identical output when reprocessing the same structure with the same settings. - Custom box vectors: Define triclinic box vectors manually instead of using standard box shapes. Specify vectors A, B, and C as comma-separated X,Y,Z components in nanometers.
- Download templates: Comma-separated residue codes to download from PDB Chemical Component Dictionary (e.g.,
ATP, GTP, HEM). Use this for non-standard residues not included in the default template library.
Understanding the results
PDB Fixer outputs corrected PDB files ready for molecular dynamics simulation. ProteinIQ emits one primary repaired PDB for each successful input and an additional summary CSV describing the whole run.
| Metric | Description |
|---|---|
| Atoms | Total atom count in the output structure |
| Residues | Number of residues (may increase if loops were added) |
| Chains | Number of protein chains retained |
| Processing applied | List of fixes performed (e.g., "Added 2,847 hydrogens, replaced 3 nonstandard residues") |
Structure viewer behavior
ProteinIQ presents batch PDB Fixer outputs as a collection of structures rather than docking poses. The viewer labels each repaired model by its source name, defaults to showing one structure at a time, and keeps the ligand-specific grouping controls hidden because PDB Fixer is not generating alternate ligand poses.
Summary file
The secondary summary records the source name, success or failure status, output filename, atom and residue counts, applied processing steps, and any error message for structures that could not be repaired. This is useful when a mixed run contains both successful and failed entries.
Validating the output
We recommend visually inspecting the fixed structure before simulation:
- Check that added loops adopt reasonable conformations (no severe clashes)
- Verify that important ligands weren't accidentally removed
- For membrane systems, confirm the protein spans the bilayer correctly
If adding missing residues produces unrealistic loop conformations, consider running energy minimization or brief MD equilibration to refine the structure. Use PDB Viewer to inspect your results directly in the browser or MolProbity for detailed geometry validation.
Best practices
Start with default settings. For standard simulation preparation, enable "Add missing heavy atoms" and "Add missing hydrogens" while keeping heterogens. This handles most common issues without aggressive modification.
Don't add missing residues unless needed. Loop modeling for large gaps (10+ residues) produces approximate conformations that require substantial equilibration. If the missing region isn't relevant to your study, leave it missing.
Remove heterogens thoughtfully. Crystallographic waters at binding sites can be important. If studying ligand binding, consider keeping all heterogens and manually curating the result.
Use appropriate ionic strength. Physiological conditions typically use 0.15 M NaCl. Zero ionic strength (pure water) can destabilize proteins. Higher concentrations (0.5–1.0 M) are appropriate for halophilic proteins or aggregation studies.
Choose the right box geometry. Dodecahedral boxes reduce computational cost by ~30% compared to cubic boxes with the same minimum padding distance. Use cubic only when your simulation software requires it.
Match water model to force field. If using AMBER14, stick with TIP3P. If using CHARMM36, TIP3P or SPC/E both work well. Mixing incompatible water models with force fields leads to incorrect thermodynamic properties.
Common workflows
Structure preparation for docking
Fix the structure with default settings, then use AutoDock Vina or DiffDock for molecular docking studies. For docking, you typically want to remove water (Remove all heterogens) but keep crystallographic ligands for reference.
MD simulation preparation
Enable all "Add" options except "Add missing residues" unless you need complete loops. Add a solvent box with appropriate ionic strength. The output is ready for equilibration in OpenMM, GROMACS, or AMBER.
Predicted structure cleanup
Structures from ESMFold, Boltz-2, or OpenFold 3 may need protonation or format adjustments. PDB Fixer can add hydrogens and standardize the structure for downstream analysis.
Frequently asked questions
Is PDB Fixer free?
Yes. ProteinIQ provides PDB Fixer online at no cost with a free account. The underlying OpenMM PDBFixer library is open-source software licensed under MIT/LGPL.
How long does PDB Fixer take to process a structure?
Processing time depends on structure size and selected options. Small proteins (under 5,000 atoms) with default settings complete in under a minute. Structures larger than 10,000 atoms are automatically routed to GPU hardware, which accelerates the energy minimization step. Adding missing residues or solvent boxes increases processing time regardless of hardware — expect several minutes for structures with extensive loop building or large solvent boxes.
Why are my missing loops in the wrong conformation?
PDB Fixer builds loops using idealized geometry followed by brief energy minimization. This provides reasonable starting conformations, but long loops (10+ residues) often need additional refinement. Run extended energy minimization or a short MD equilibration to relax the structure.
What's the difference between PDB Fixer and PROPKA?
PDB Fixer adds missing atoms and prepares structures for simulation. PROPKA predicts pKa values for titratable residues without modifying the structure. Use PROPKA first to determine appropriate pH, then run PDB Fixer with that pH value to add hydrogens with correct protonation states.
Can I use PDB Fixer output directly for MD simulations?
Yes, with appropriate settings. Enable "Add solvent box" to create a solvated system with counterions. The output PDB includes periodic box vectors and is ready for equilibration in OpenMM, GROMACS, or AMBER.
Why did PDB Fixer remove my ligand?
Check your "Heterogens" setting. If set to Remove all, all non-protein molecules including ligands are deleted. Use Keep all to preserve ligands and crystallographic waters.
How do I fix a structure with nonstandard amino acids?
Enable "Replace nonstandard residues" to convert modified residues (selenomethionine, phosphoserine, etc.) to their standard equivalents. If you need to keep modifications, disable this option and ensure the residue templates are available—use "Download templates" to fetch specific residue definitions from the PDB Chemical Component Dictionary.
What force field should I use?
AMBER14 works well for most protein simulations and is the recommended default. Use CHARMM36 if you plan to run simulations with CHARMM-compatible software or if you're studying nucleic acids or specific lipid systems where CHARMM parameters are preferred.
Can PDB Fixer handle multiple chains?
Yes. PDB Fixer processes all chains in the input structure. Use "Remove chains" to exclude specific chains from the output if you only need certain parts of a complex.
How do I prepare a membrane protein?
Enable "Add lipid membrane" instead of "Add solvent box." Select the appropriate lipid type (POPC for general use, POPE for bacterial membranes) and position the membrane center appropriately for your protein's transmembrane domain.