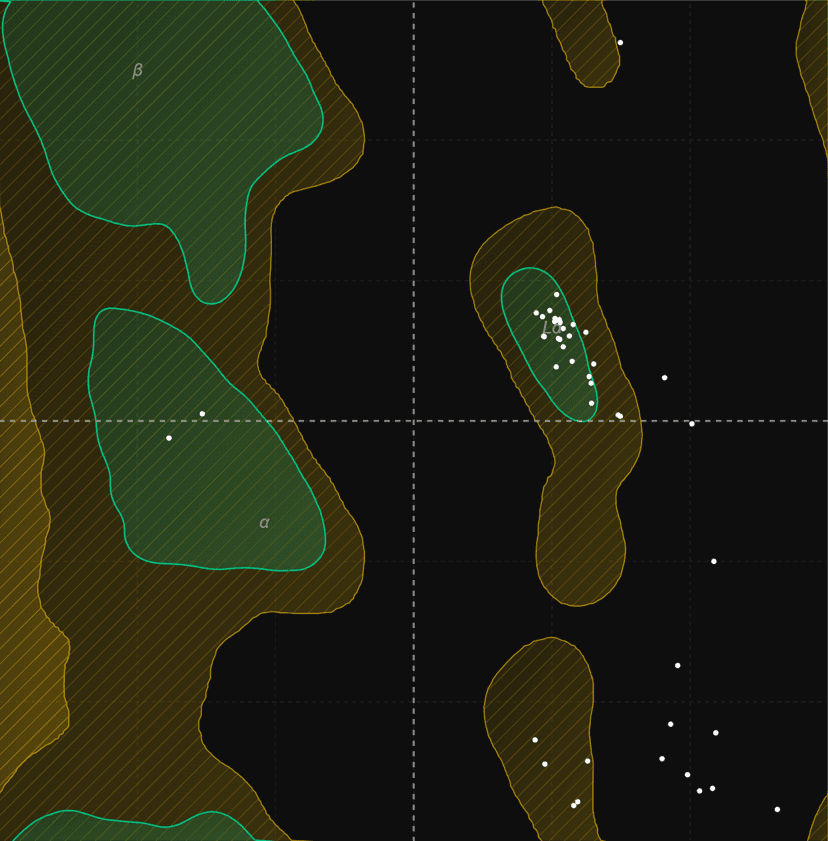
Ramachandran plot
Analyze protein backbone phi/psi angles and classify favored, allowed, and outlier residues.
Related tools

MSA Viewer
Interactive viewer for multiple sequence alignments with color-coded residues and consensus sequence
MolProbity
Validate protein structure quality with all-atom contact analysis, Ramachandran plots, rotamer assessment, and geometry checks.
DockQ
Assess docking model quality by comparing predicted complexes against native references. DockQ v2.1.3 supports protein, nucleic-acid, and supported small-molecule interfaces with faithful upstream metrics.
IPSAE
Scoring function for interprotein interactions in AlphaFold2, AlphaFold3 and Boltz predictions. Calculates ipSAE, ipTM, pDockQ, pDockQ2, and LIS scores to assess protein-protein interface quality.
PoseBusters
PoseBusters validates generated or docked molecular poses with chemically and structurally grounded quality checks for molecular geometry, intermolecular interactions, and optional reference-pose agreement.

PDB viewer
Visualize and analyze protein structures in 3D using Mol* (molstar), the same viewer used by AlphaFold DB and RCSB PDB

Protein charge plot
Plot net charge vs pH for protein sequences. Visualize how protein charge changes across pH 0-14 and identify the isoelectric point (pI) where the net charge crosses zero.
Hydropathy plot
Generate Kyte-Doolittle hydropathy plots to visualize hydrophobic and hydrophilic regions along protein sequences. Identify transmembrane domains and surface-exposed regions.
Hydrophobicity plot
Generate hydrophobicity plots using 24 different amino acid scales. Visualize hydrophobic and hydrophilic regions for protein analysis, epitope prediction, and membrane protein studies.

Protein scale profiler
Generate amino acid property profiles using 42 different scales spanning hydrophobicity, secondary structure propensity, flexibility, polarity, surface accessibility, antigenicity, and more.
What is a Ramachandran plot?
The Ramachandran plot is a two-dimensional visualization of backbone dihedral angles in proteins. Developed in 1963 by G.N. Ramachandran, C. Ramakrishnan, and V. Sasisekharan, it maps the phi (φ) angle against the psi (ψ) angle for each amino acid residue in a protein structure.
The plot reveals which backbone conformations are sterically possible by showing favored, allowed, and outlier regions. Certain angle combinations cause atoms to clash, making those conformations energetically unfavorable. This creates characteristic "allowed" regions that correspond to common secondary structures.
Ramachandran analysis is a standard quality metric in structural biology. High-resolution crystal structures typically show >98% of residues in favored regions. Deviations often indicate errors in structure determination or unusual but functionally important conformations.
How does a Ramachandran plot work?
Backbone dihedral angles
The protein backbone contains three types of dihedral angles. The phi (φ) angle describes rotation around the N-Cα bond, while the psi (ψ) angle describes rotation around the Cα-C bond. The omega (ω) angle at the peptide bond is typically fixed at 180° due to partial double-bond character.
The phi angle is calculated from four atoms: C(i-1), N(i), Cα(i), and C(i). The psi angle requires N(i), Cα(i), C(i), and N(i+1). This dependency on neighboring residues means terminal residues cannot have both angles calculated.
Steric constraints
Ramachandran's original analysis treated atoms as hard spheres with van der Waals radii. Systematic sampling of all possible φ/ψ combinations revealed that most are sterically forbidden due to atomic clashes. Only specific angle ranges allow the backbone atoms to avoid collision.
The resulting allowed regions correspond directly to protein secondary structures. Alpha helices cluster near φ = -60° and ψ = -45°. Beta sheets occupy the region around φ = -120° and ψ = +120°. Left-handed helices appear in a smaller region with positive phi values.
Residue-specific regions
Different amino acids have distinct allowed regions based on their side chain properties:
- General residues: Most amino acids follow the standard Ramachandran distribution with restricted phi values (predominantly negative)
- Glycine: Lacks a side chain, allowing much greater conformational freedom with nearly symmetric allowed regions
- Proline: The cyclic side chain restricts phi to approximately -60°, severely limiting allowed conformations
- Pre-proline: Residues preceding proline show altered distributions due to steric interactions with the proline ring
Region classification
Modern Ramachandran analysis uses empirical boundaries derived from high-resolution structures. This tool implements the classification system from Lovell et al. (2003), based on statistical analysis of ~500 high-quality crystal structures:
- Favored: Contains ~98% of residues in well-refined structures. Indicates optimal backbone geometry.
- Allowed: Contains an additional ~2% of residues. Represents conformations that are sterically possible but less common.
- Outlier: Residues outside allowed regions. May indicate modeling errors or unusual but real conformations.
Input requirements
- Protein structure: A PDB file containing atomic coordinates. The tool requires complete backbone atoms (N, Cα, C) for angle calculation. PDB IDs can be entered directly for automatic fetching from the RCSB database.
Understanding the results
Plot interpretation
Each point on the plot represents one residue's φ/ψ angles. Green shaded regions indicate favored conformations; yellow regions show allowed conformations. Points outside colored regions are outliers.
Hover over points to see residue details including name, number, chain, exact angles, and region classification. Use the highlight controls to filter by residue type (general, glycine, proline, pre-proline) or to show only outliers.
Statistics panel
The statistics summarize structure quality:
- >98% favored: Expected for high-quality structures
- 95-98% favored: Acceptable for most purposes
- <95% favored: May indicate structural problems requiring investigation
Outlier analysis
Outliers are listed in a dedicated table with their exact angles. Not all outliers indicate errors. Functionally important residues sometimes adopt strained conformations for catalysis or binding. Compare outlier positions with functional annotations before concluding they represent errors.
Common applications
Ramachandran analysis serves multiple purposes in structural biology:
- Model validation: Assess quality of crystal structures, NMR ensembles, or homology models
- Structure refinement: Identify residues requiring manual adjustment during model building
- Comparative analysis: Evaluate structural changes between conformational states or mutants
- Education: Understand the relationship between backbone geometry and secondary structure
Cost
Ramachandran plot generation is free and runs entirely in your browser. No credits required.