Related tools
ESM-2
ESM-2 is a 650M parameter protein language model from Meta AI trained on 250M protein sequences. Generate rich sequence representations for downstream tasks like structure prediction, function annotation, and variant effect prediction.
ProstT5
ProstT5 is a protein language model that bidirectionally translates between amino acid sequences and 3Di structural tokens. It enables fast structure-based searches and inverse folding by encoding structural information into a sequence-like representation.
AbLang
Restore missing residues in antibody sequences using a language model trained on the Observed Antibody Space (OAS) database. Achieves better restoration than IMGT germlines or ESM-1b while being 7x faster.
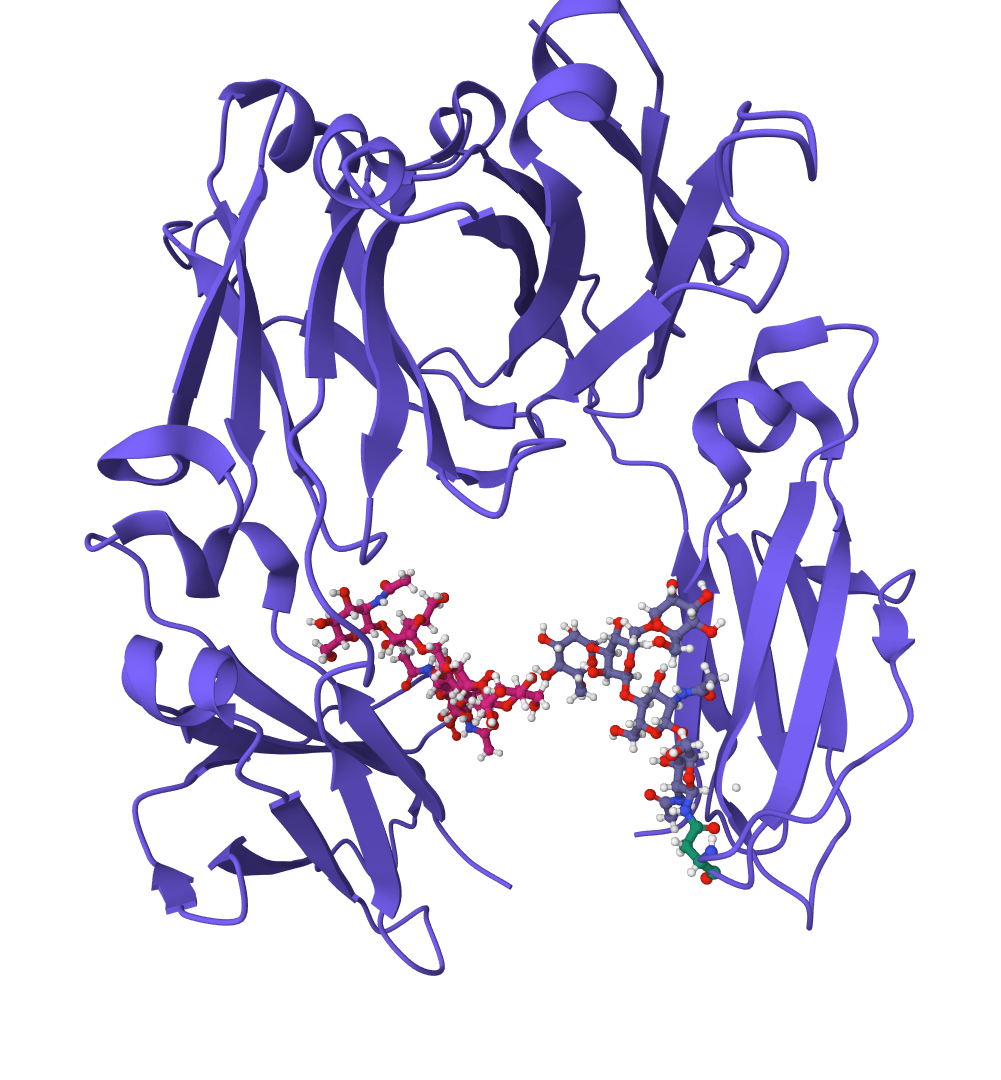
ScanNet
Geometric deep learning model for predicting protein binding sites directly from 3D structure. Identifies where proteins interact with other proteins, antibodies, or disordered proteins with high accuracy, including for novel protein folds.

DR-BERT
DR-BERT is a compact protein language model that predicts intrinsically disordered regions (IDRs) in proteins. It outputs per-residue disorder probability scores (0–1) from amino acid sequences, enabling fast and accurate annotation of disordered regions without structural data.
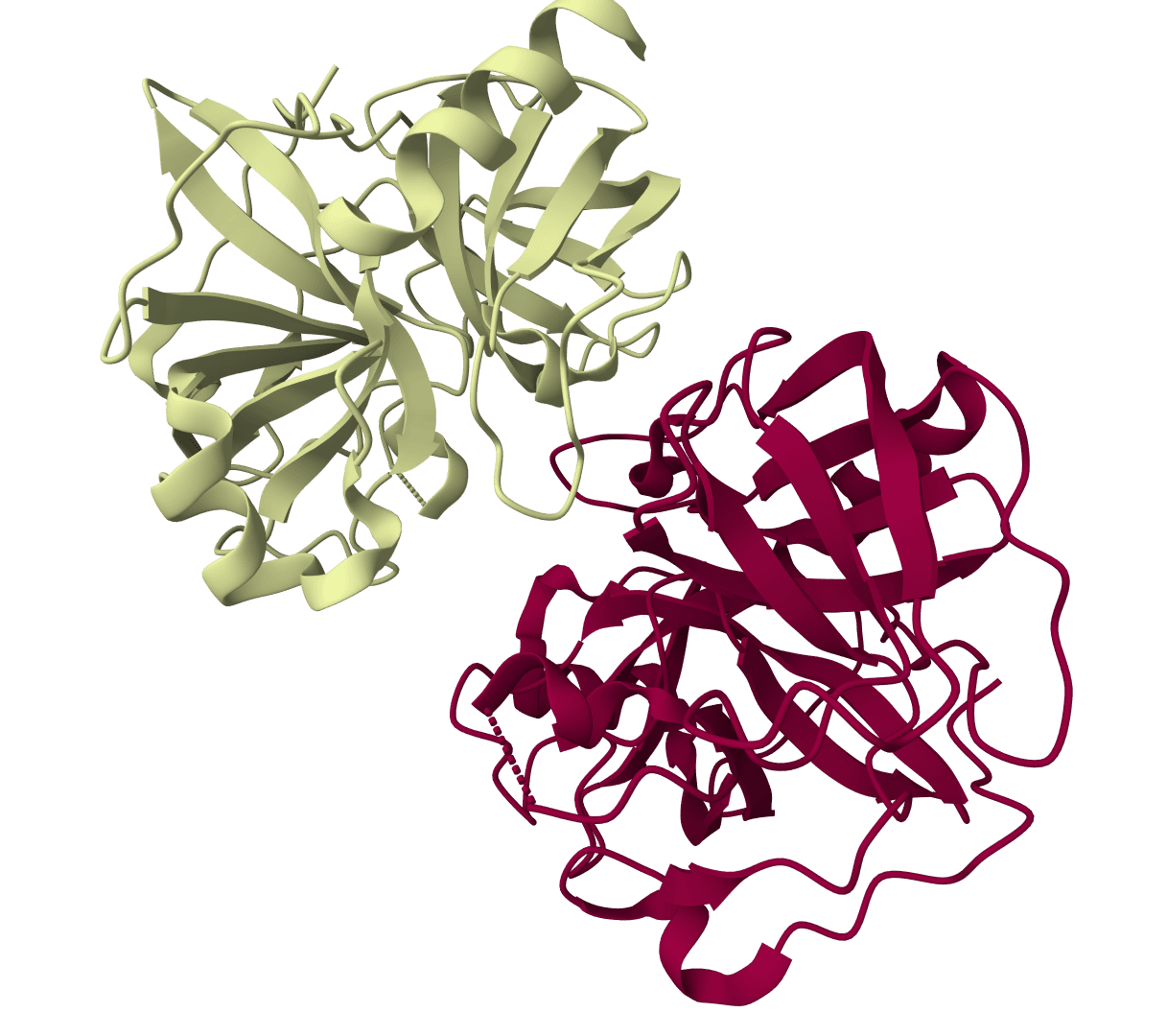
AF-Cluster
Cluster Multiple Sequence Alignments to predict alternative protein conformations with AlphaFold2. Uses DBSCAN clustering to identify sequence subgroups.
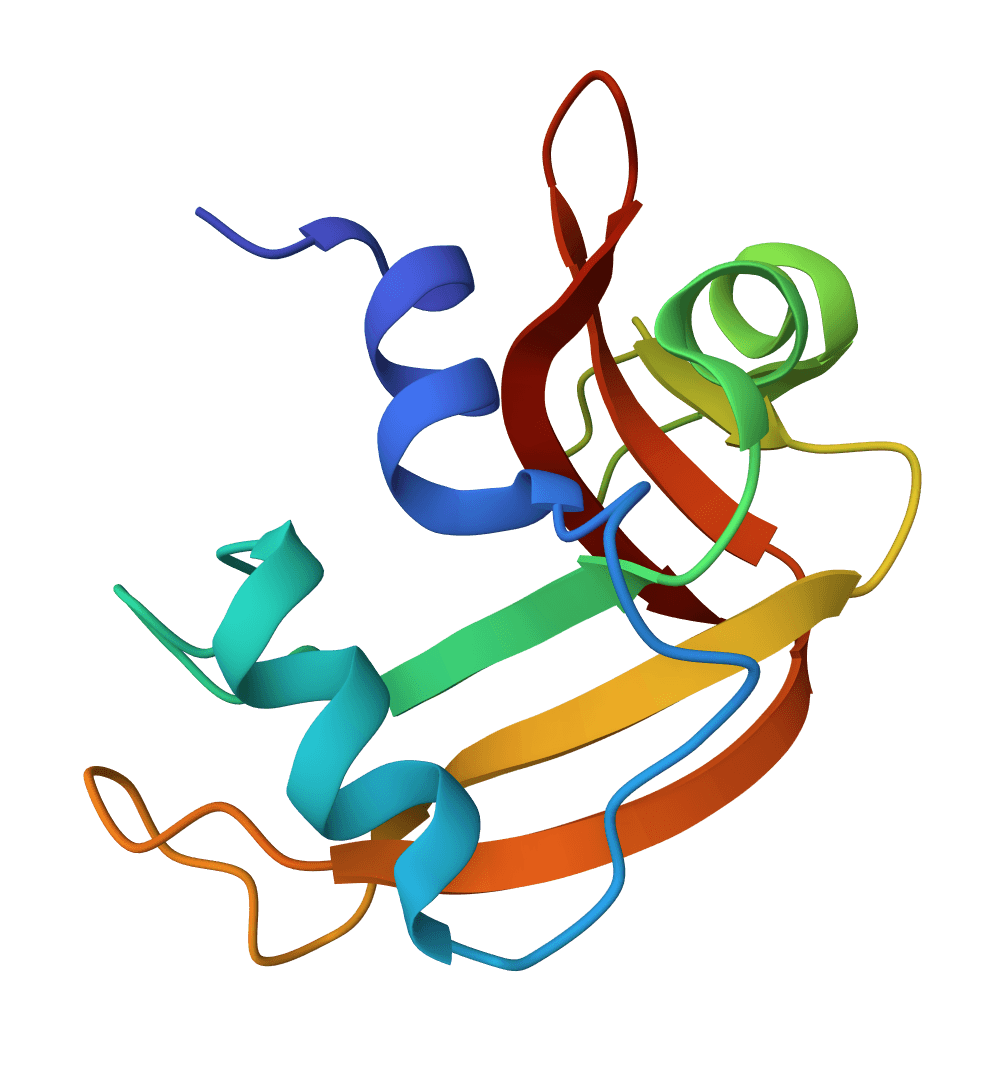
ANARCI
Number antibody and T cell receptor variable domain sequences using multiple numbering schemes (IMGT, Chothia, Kabat, Martin, AHo, Wolfguy). Identifies chain type, species, and assigns germline genes.
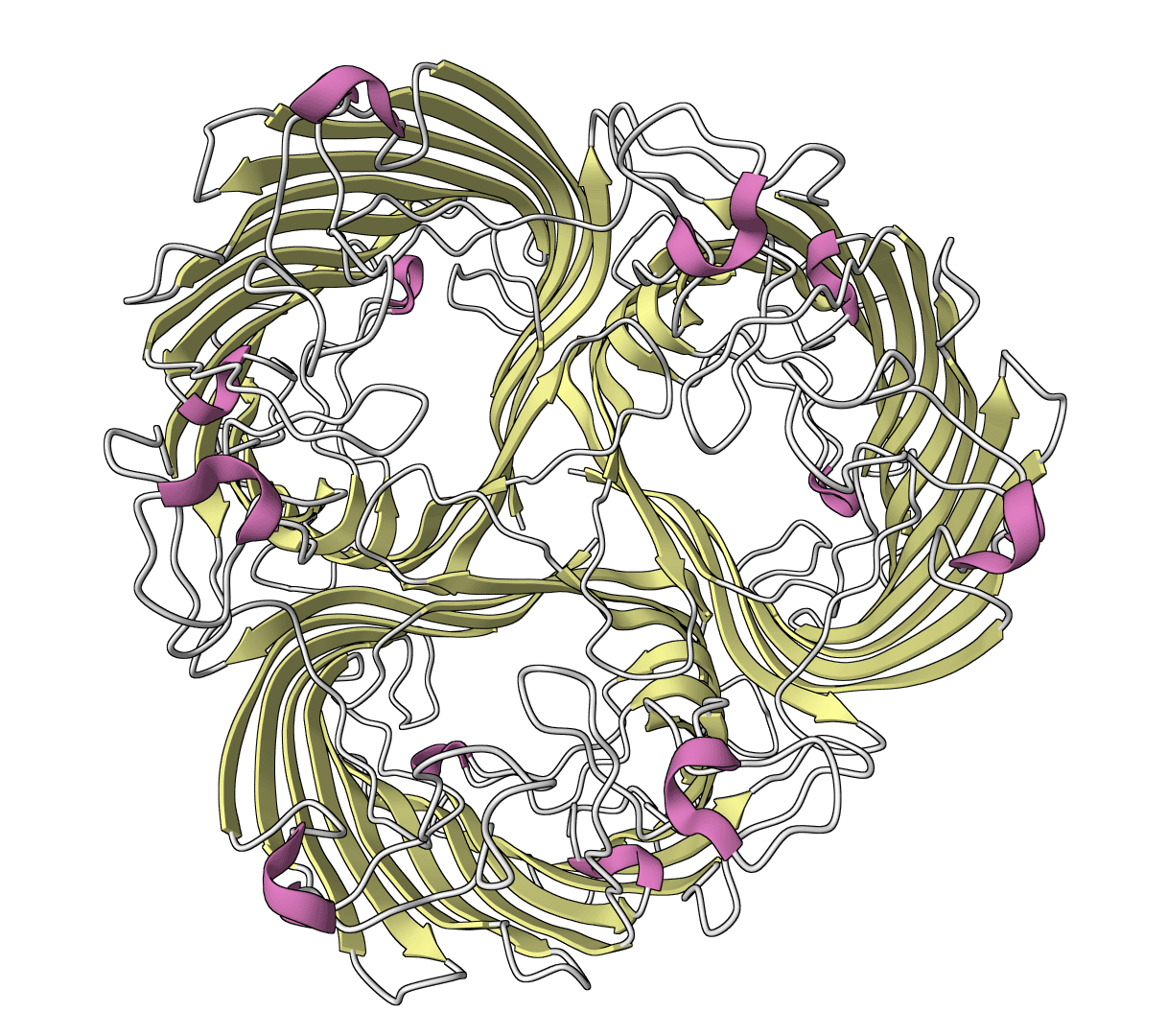
CleaveNet
Official CleaveNet wrapper for matrix metalloproteinase cleavage prediction and peptide generation. Predict cleavage z-scores plus uncertainty across 17 MMP variants, evaluate against truth z-scores, or generate candidate peptides unconditionally or from MMP z-score profiles.
Humatch
Humatch is an antibody humanization tool that transforms non-human antibody sequences into humanized variants. Uses three lightweight CNNs to identify optimal human V-genes and generate paired heavy and light chain sequences with minimal edits while maintaining functionality.
IgBLAST
Analyze immunoglobulin (antibody) and T cell receptor variable domain sequences. Identifies V/D/J gene segments, delineates CDR regions, and analyzes rearrangement junctions.
What is AbLang-2?
AbLang-2 is an antibody-specific language model designed to predict non-germline (NGL) residues in antibody sequences. Developed by Tobias Olsen and colleagues at the Oxford Protein Informatics Group, it addresses a fundamental limitation of earlier antibody language models: their tendency to predict germline residues while ignoring the somatic mutations that make antibodies functional.
Antibody sequences are heavily biased toward their germline templates. During somatic hypermutation, only a small fraction of residues mutate to create binding specificity against a target. When trained on natural antibody repertoires, models like AbLang-1, AntiBERTy, and Sapiens learn to predict germline residues with overwhelming probability (84-87%) because that's what most residues are.
How does AbLang-2 work?
The model architecture consists of 12 transformer layers with 480-dimensional embeddings (45 million parameters). Training proceeds in two phases: pre-training on 35.6 million unpaired VH and VL sequences from the Observed Antibody Space (OAS) database, followed by fine-tuning on 1.26 million paired heavy/light chain sequences.
The key innovation is the use of focal loss, which down-weights the loss contribution from well-predicted labels. Since germline residues are easy to predict, focal loss reduces their influence on the gradient and forces the model to learn NGL patterns. This improved NGL perplexity from the 14-39 range seen in prior models down to 10-12, achieving 15% cumulative probability for known NGL residues compared to less than 2% for earlier models.
AbLang-2 outputs two types of representations:
- Pseudo-likelihoods: A score indicating how well each position matches the learned antibody distribution. Lower values suggest unusual or potentially problematic residues.
- Embeddings: 480-dimensional vectors encoding the sequence context, suitable as features for downstream machine learning tasks.
How to use AbLang-2 online
ProteinIQ provides browser-based access to AbLang-2 with no software installation or command-line usage required. Any registered user can run the model directly from the web interface.
Input
| Input | Description |
|---|---|
Antibody sequences | FASTA format. For paired mode, join heavy and light chains with a pipe: VH_sequence|VL_sequence. For unpaired mode, provide individual chains with descriptive headers. |
Settings
| Setting | Description |
|---|---|
Sequence mode | Paired (VH|VL) expects pipe-separated heavy and light chains. Unpaired processes individual chains separately. |
Output type | Pseudo-likelihoods returns per-sequence scores. Sequence embeddings returns 480-dimensional vectors. Both returns both outputs. |
Output
Results appear as a spreadsheet with columns depending on the selected output type:
| Column | Description |
|---|---|
Sequence ID | Identifier from FASTA header |
Sequence | Input sequence |
Length | Number of residues |
Pseudo-likelihood | Score reflecting how well the sequence fits the antibody language model |
Embedding dim | Dimensionality of the embedding vector (480) |
When downloading results with embeddings, the full 480-dimensional vectors are included in the exported file.
Interpreting pseudo-likelihoods
Pseudo-likelihood scores represent how "antibody-like" a sequence appears to the model. Higher scores indicate sequences that better match the statistical patterns learned from natural antibodies. There is no universal threshold for good or bad scores, but relative comparisons are meaningful:
- Sequences with unusually low scores may contain problematic residues or frameshifts
- Comparing variants of the same antibody can identify mutations that disrupt antibody-like properties
- Tracking scores across humanization or optimization rounds can guide design decisions
Limitations
AbLang-2 excels at predicting NGL residues but does not guarantee that suggested mutations will improve binding or stability. The model reflects statistical patterns in natural antibodies, not experimental binding data.
Paired mode requires both chains. If only one chain is available, use unpaired mode with the appropriate chain type.