Related tools
MAFFT
Perform multiple sequence alignment using MAFFT (Multiple Alignment using Fast Fourier Transform). Supports multiple algorithms from fast progressive to highly accurate iterative methods.
MUSCLE5
Perform multiple sequence alignment using MUSCLE5 (MUltiple Sequence Comparison by Log-Expectation). Uses the PPP algorithm for high-quality alignments with support for ensemble generation.
HMMER
Sensitive sequence homology search using profile hidden Markov models. More accurate than BLAST for detecting remote homologs, ideal for finding evolutionarily distant protein family members.
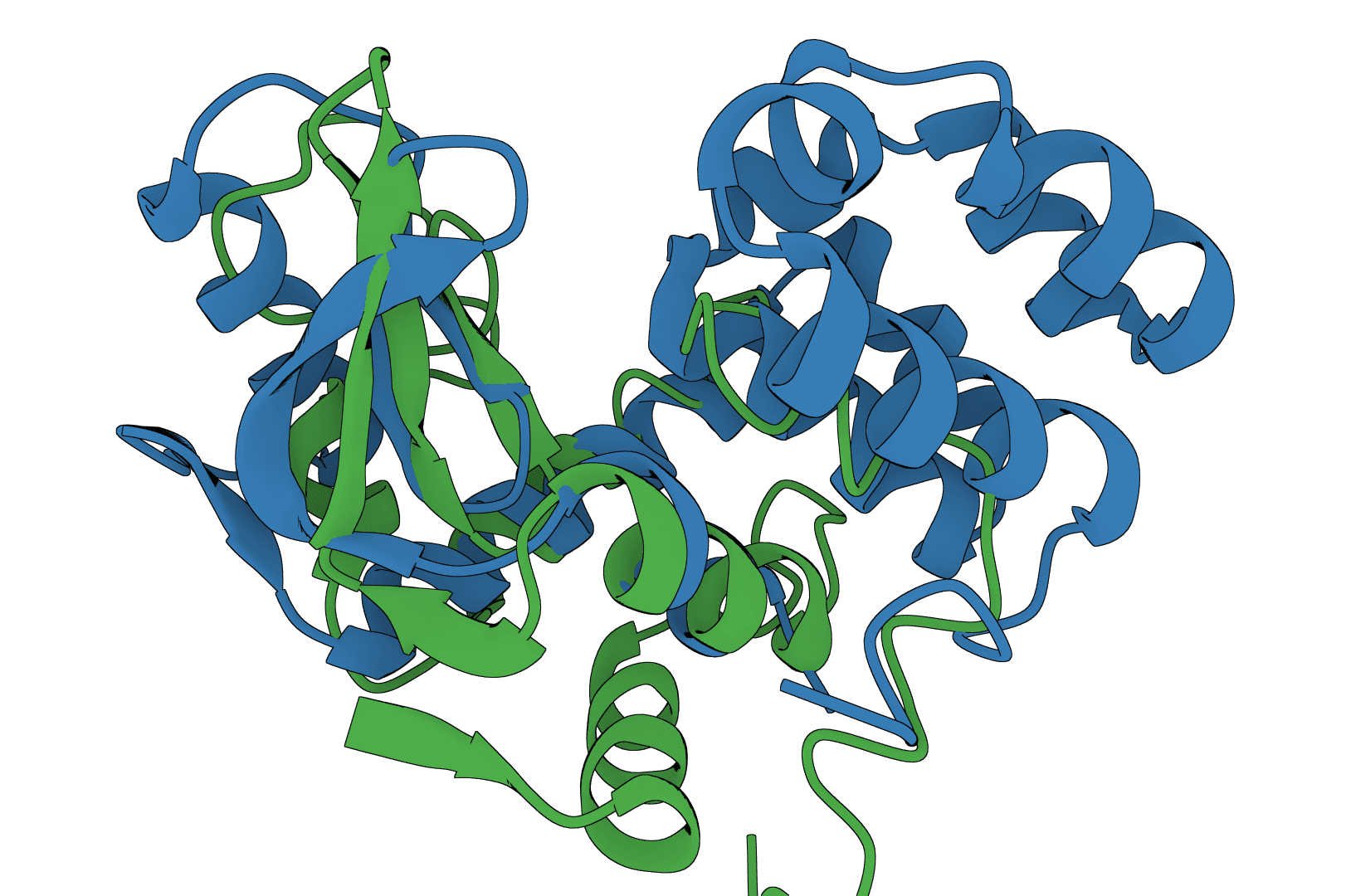
USAlign
USAlign (Universal Structure Alignment) aligns protein, RNA, and DNA structures to compute TM-scores and generate superposed structures. Compare 3D structures to assess structural similarity.
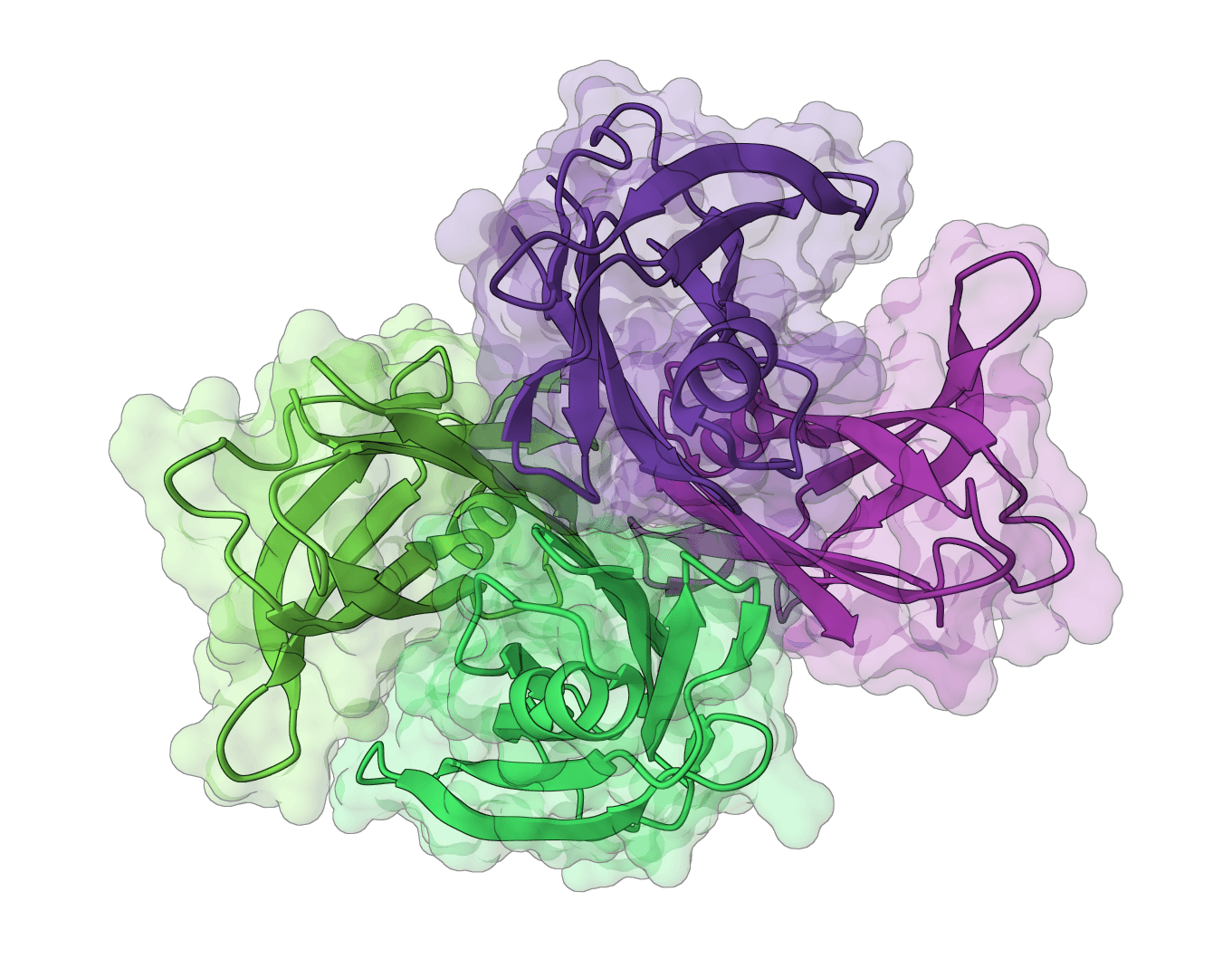
IgBLAST
Analyze immunoglobulin (antibody) and T cell receptor variable domain sequences. Identifies V/D/J gene segments, delineates CDR regions, and analyzes rearrangement junctions.
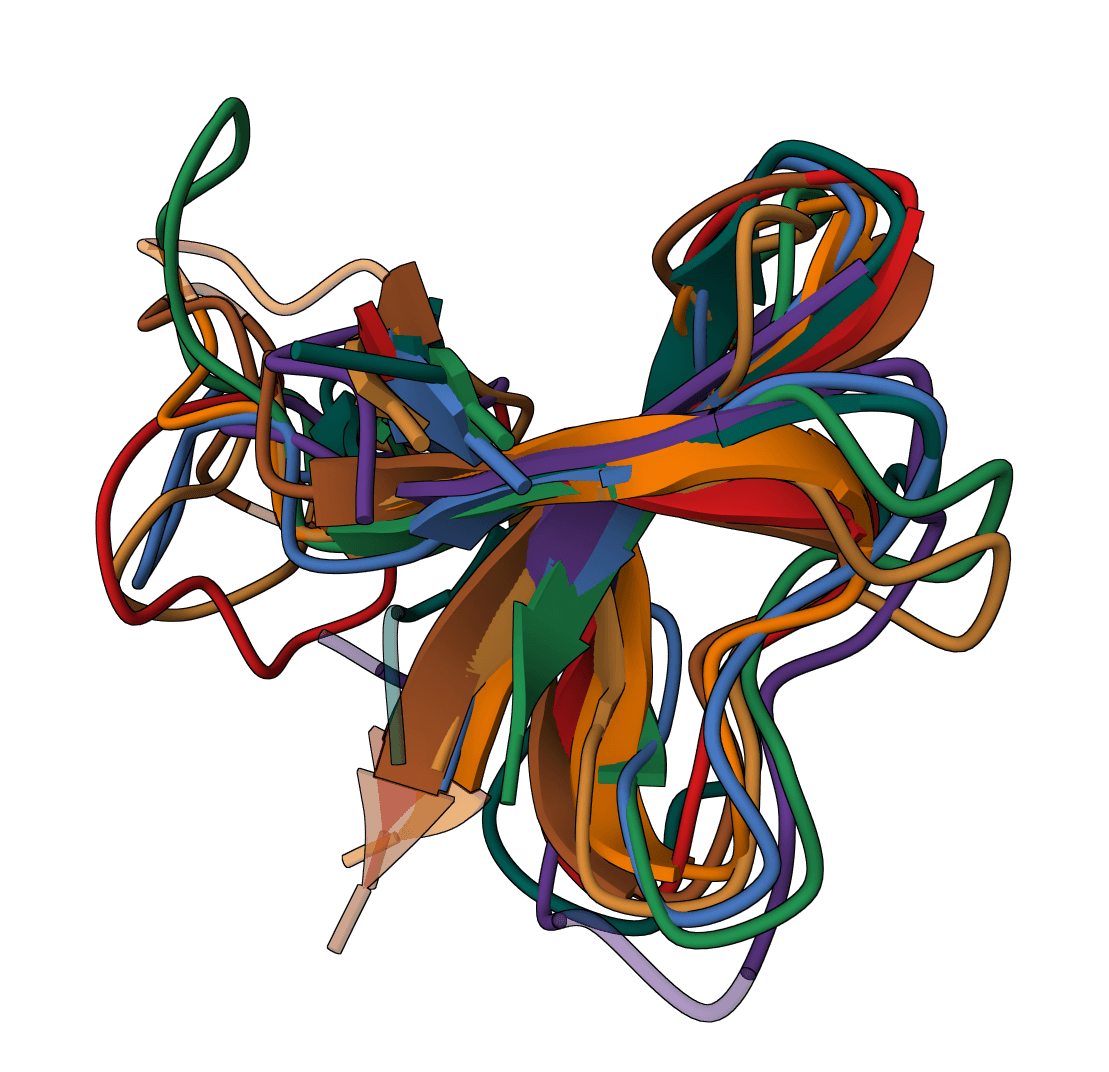
Clustal Omega
Perform multiple sequence alignment on protein or nucleotide sequences using the Clustal Omega algorithm.
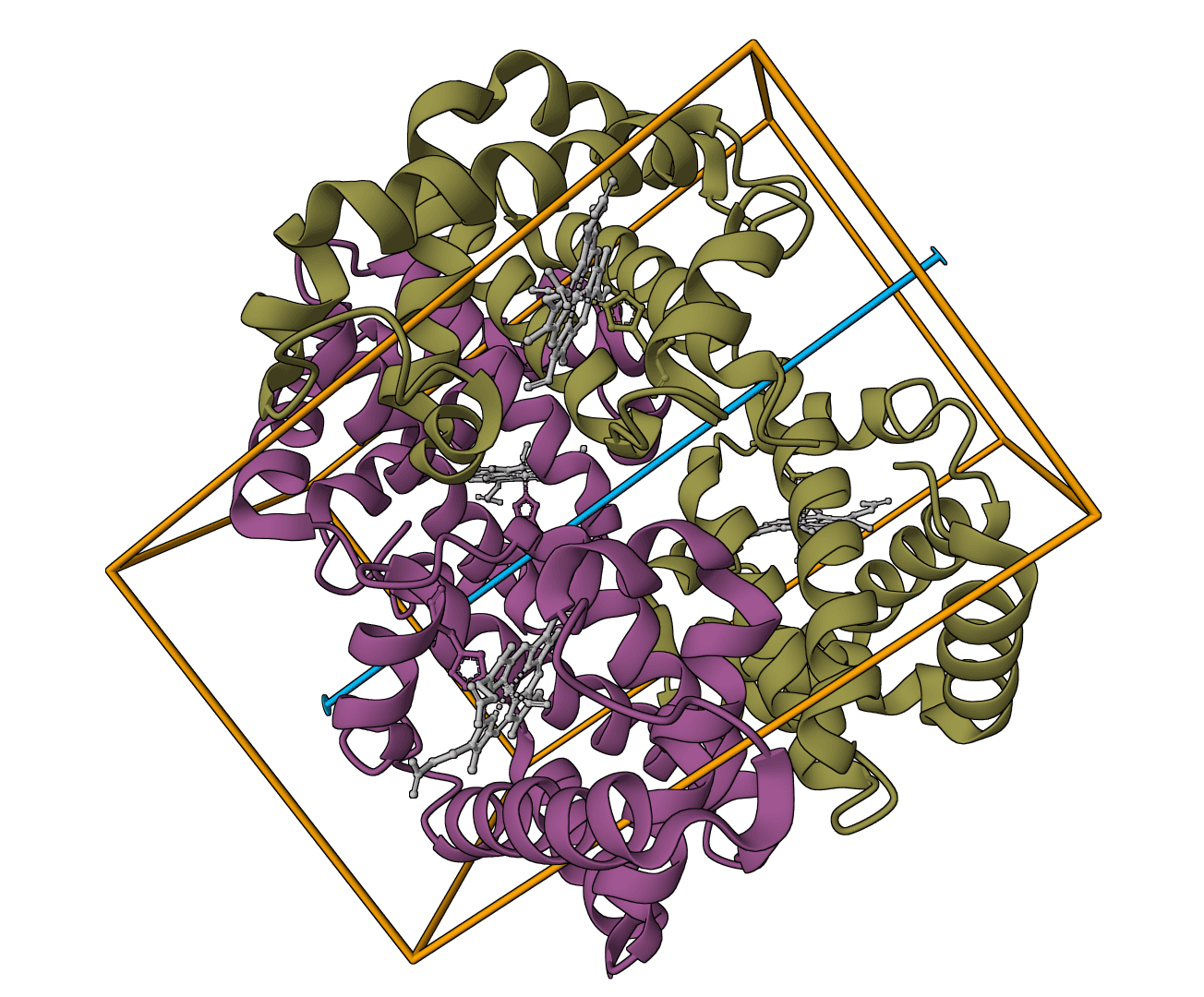
FastTree
Infer approximately-maximum-likelihood phylogenetic trees from alignments of nucleotide or protein sequences.
IQ-TREE
Build phylogenetic trees using maximum likelihood with automatic model selection (ModelFinder) and ultrafast bootstrap support.
ANARCI
Number antibody and T cell receptor variable domain sequences using multiple numbering schemes (IMGT, Chothia, Kabat, Martin, AHo, Wolfguy). Identifies chain type, species, and assigns germline genes.
MUMmer4
Rapidly align and compare DNA sequences using MUMmer4 nucmer. Perform pairwise genome comparisons to identify SNPs, indels, and structural variants between reference and query genomes.
What is MMseqs2?
MMseqs2 (Many-against-Many sequence searching) is an ultra-fast tool for searching and clustering protein and nucleotide sequences. It runs up to 10,000 times faster than BLAST while maintaining comparable sensitivity, making it practical to search millions of sequences in minutes rather than days.
The tool serves two primary use cases: finding homologous sequences in a database (search mode) and grouping similar sequences together (clustering mode). For researchers working with large datasets from metagenomics, genomics, or protein family analysis, MMseqs2 enables analyses that would be computationally infeasible with traditional tools.
For multiple sequence alignment after identifying homologs, see Clustal Omega. For phylogenetic tree construction, use FastTree.
How does MMseqs2 work?
MMseqs2 achieves its speed through a two-stage filtering approach that eliminates unrelated sequences before performing expensive alignments.
K-mer prefiltering
The first stage uses k-mer matching to rapidly identify candidate sequences. MMseqs2 extracts short sequence fragments (k-mers) and stores them in a memory-based index. For each query, it generates lists of similar k-mers and looks for "double consecutive k-mer matches"—two similar k-mers appearing on the same diagonal in a sequence alignment.
This prefiltering rejects approximately 99.99% of sequences that have no meaningful similarity, dramatically reducing the computational burden.
Smith-Waterman alignment
Sequences passing prefiltering undergo vectorized Smith-Waterman alignment, the gold-standard local alignment algorithm. This stage calculates precise alignment scores, sequence identity, and E-values for the final results.
Sensitivity parameter
The sensitivity parameter (-s) controls how many similar k-mers are considered during prefiltering. Higher values (up to 7) consider more distant k-mer variants, finding more remote homologs at the cost of speed:
1-4: Fast searches, finds close homologs5-6: Balanced, suitable for most applications7: Maximum sensitivity, comparable to BLAST
Operating modes
MMseqs2 offers three distinct modes optimized for different tasks.
Search mode
Compares query sequences against a target database to find homologs. Outputs BLAST-compatible tabular format (m8) with 12 columns: query ID, target ID, sequence identity, alignment length, mismatches, gap openings, query/target start/end positions, E-value, and bit score.
Cluster mode
Groups similar sequences using a greedy set-cover algorithm. Each cluster has a representative sequence, and all other members are similar to the representative above the specified thresholds. The output shows representative-member pairs.
Linclust mode
A linear-time clustering algorithm for very large datasets (millions of sequences). Linclust sacrifices some clustering quality for speed, scaling linearly with database size rather than quadratically.
Clustering parameters
When using cluster or linclust mode, several parameters control how sequences are grouped.
Minimum sequence identity
Sets the similarity threshold for clustering. Sequences must share at least this fraction of identical residues to be grouped together. Common thresholds:
0.9(90%): Near-identical sequences, removes redundancy0.5(50%): Same protein family0.3(30%): Remote homologs, twilight zone of sequence similarity
Coverage modes
Coverage defines how much of each sequence must align. The coverage mode determines which sequence lengths are considered:
- Mode 0: Coverage of both query and target—alignment must cover a fraction of the longer sequence
- Mode 1: Coverage of target only
- Mode 2: Coverage of query only
- Mode 3: Bidirectional—both sequences must meet the coverage threshold independently
Clustering algorithms
Four algorithms are available for grouping sequences:
- Set-cover (mode 0): Recommended default. Greedily selects cluster representatives that cover the most sequences.
- Connected component (mode 1): Groups all transitively connected sequences. Produces larger, more inclusive clusters.
- Greedy sequential (mode 2): Processes sequences in input order, similar to CD-HIT.
- Greedy by length (mode 3): Selects longest sequences as representatives first.
Search parameters
E-value threshold
The expectation value represents the number of hits expected by chance in a database of this size. Lower E-values indicate more significant matches:
0.001: Stringent, high-confidence homologs only0.01: Standard threshold1-10: Permissive, includes weak similarities
Maximum hits
Limits the number of target sequences reported per query. Useful for controlling output size when searching large databases.
Understanding the results
Search output
Results appear in BLAST m8 tabular format:
| Column | Description |
|---|---|
| query | Query sequence identifier |
| target | Target sequence identifier |
| pident | Percentage sequence identity |
| alnlen | Alignment length |
| mismatch | Number of mismatches |
| gapopen | Number of gap openings |
| qstart/qend | Query alignment coordinates |
| tstart/tend | Target alignment coordinates |
| evalue | E-value (significance) |
| bits | Bit score |
Clustering output
Clustering produces a two-column table:
| Column | Description |
|---|---|
| representative | Cluster representative sequence ID |
| member | Sequence belonging to this cluster |
When representative equals member, that sequence is either a singleton (no similar sequences found) or the cluster representative itself.
Best practices
For database searches, start with the default sensitivity (5.7) and adjust based on results. If you're missing expected homologs, increase sensitivity. If searches are too slow, decrease it.
For clustering, choose thresholds based on your biological question. Redundancy removal typically uses 90-95% identity. Protein family clustering works well at 30-50% identity.
Use linclust instead of cluster when processing more than 100,000 sequences—the speed difference becomes substantial at scale.