Related tools

DR-BERT
DR-BERT is a compact protein language model that predicts intrinsically disordered regions (IDRs) in proteins. It outputs per-residue disorder probability scores (0–1) from amino acid sequences, enabling fast and accurate annotation of disordered regions without structural data.
CleaveNet
Official CleaveNet wrapper for matrix metalloproteinase cleavage prediction and peptide generation. Predict cleavage z-scores plus uncertainty across 17 MMP variants, evaluate against truth z-scores, or generate candidate peptides unconditionally or from MMP z-score profiles.

CpG Island Finder
Identify CpG islands in DNA sequences using the Gardiner-Garden and Frommer criteria. Analyze GC content, CpG density, and observed/expected ratios.
GC content calculator
Calculate GC content and nucleotide composition of DNA/RNA sequences. Analyze individual sequences or get combined statistics.

MSA Viewer
Interactive viewer for multiple sequence alignments with color-coded residues and consensus sequence
ORF Finder
Find all Open Reading Frames (ORFs) in DNA sequences. Searches all six reading frames and supports multiple genetic codes.

RAxML-NG
Perform maximum-likelihood phylogenetic tree inference with RAxML-NG for aligned protein or DNA sequences. Supports ML search, bootstrap analysis, and upstream-style automatic model-family selection.
SuperWater
Predict protein hydration sites from a structure using a diffusion model with ESM features and a confidence-filtering head.
Aggrescan3D
Faithful static-mode Aggrescan3D wrapper for per-residue aggregation propensity analysis from a single protein structure.
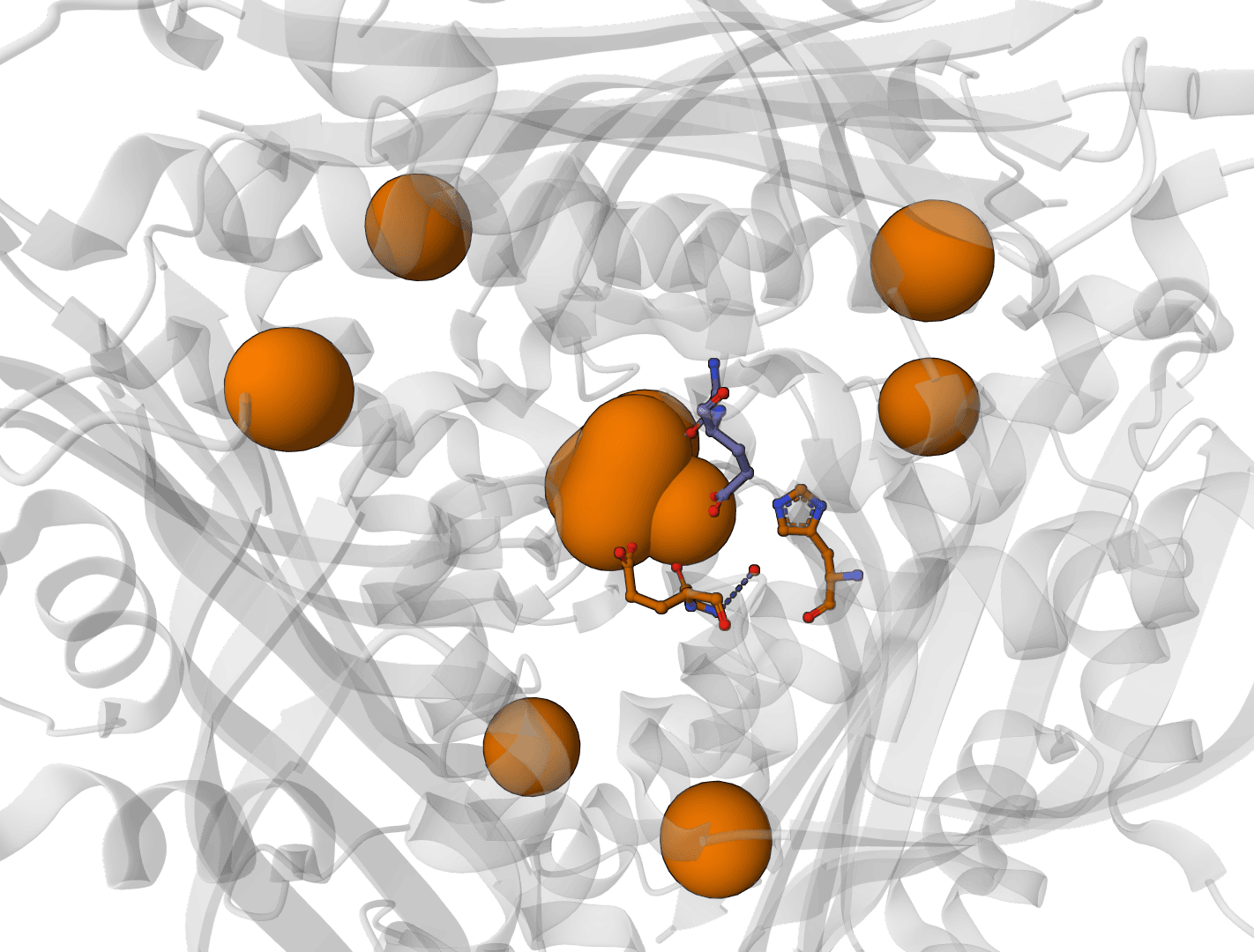
AllMetal3D
Predict metal and water binding sites in protein structures using 3D convolutional neural networks (AllMetal3D + Water3D).
What is AlphaGenome?
AlphaGenome is a deep learning model from Google DeepMind that predicts how genetic variants affect gene regulation. Given a genomic position and a variant (reference → alternate allele), AlphaGenome computes expression changes across 667 cell types and tissues at single base-pair resolution.
The model processes up to 1 million base pairs of DNA sequence and outputs predictions for gene expression (RNA-seq), splicing patterns, chromatin accessibility, and other regulatory features. Unlike models that focus on protein-coding regions (about 2% of the genome), AlphaGenome specializes in the non-coding 98% where most disease-associated variants reside.
AlphaGenome builds on DeepMind's earlier Enformer model and complements AlphaMissense, which predicts effects of variants within protein-coding regions.
How to use AlphaGenome online
ProteinIQ provides a browser interface to AlphaGenome, handling the API communication and result formatting without requiring local Python setup.
AlphaGenome on ProteinIQ uses a Bring Your Own Key model. This means the tool requires a personal API key from Google DeepMind rather than using ProteinIQ's infrastructure.
Why do you need your own key?
AlphaGenome is licensed by DeepMind for non-commercial research use only. This means ProteinIQ cannot host the model on its own servers or provide a shared API key. Doing so would violate the commercial use restrictions. By using a personal API key, researchers access AlphaGenome directly through DeepMind's infrastructure under their own non-commercial research agreement. Users are responsible for ensuring their use complies with DeepMind's Terms of Use.
How to get an API key
- Visit deepmind.google.com/science/alphagenome
- Sign in with a Google account
- Accept the Terms of Use (non-commercial research only)
- Copy the generated API key
The API key is free for academic and non-commercial research. Organizations seeking commercial access should contact DeepMind through their dedicated inquiry form.
API key security
ProteinIQ does not store API keys in its database. Keys are stripped from job records before storage and are only used transiently to communicate with DeepMind's API. However, the key does pass through ProteinIQ's processing servers during job execution.
Inputs
| Field | Description |
|---|---|
AlphaGenome API Key | Personal API key from DeepMind (required) |
Job name | Label for identifying the job in history |
Settings
| Setting | Description |
|---|---|
Chromosome | Chromosome identifier (e.g., chr1, chr22, chrX) |
Variant position | Genomic coordinate of the variant. The analysis window centers on this position. |
Reference bases | Original nucleotide(s) at the position (e.g., A, GT) |
Alternate bases | Variant nucleotide(s) to compare (e.g., C, GAA) |
Window size | Analysis region: 16K bp (fastest), 128K bp, or 512K bp. Larger windows capture more regulatory context but produce larger output files. |
Output settings
| Setting | Description |
|---|---|
Requested outputs | Prediction type. Currently supports RNA-seq (gene expression). |
Ontology terms | Optional comma-separated UBERON (tissue) or CL (cell type) codes for tissue-specific predictions. Example: UBERON:0002107 for liver, CL:0000057 for fibroblast. |
Results
The output includes:
- Summary statistics: Mean, max, min, and standard deviation of expression values for reference and alternate alleles, plus the predicted effect (difference)
- Full resolution data: Downloadable
.npzfile containing expression predictions at every position in the window across all 667 cell types
| Metric | Interpretation |
|---|---|
| Effect Mean | Average expression change across all positions and cell types |
| Effect at Variant | Predicted expression change at the exact variant position |
| Positive effect | Variant increases expression relative to reference |
| Negative effect | Variant decreases expression relative to reference |
How AlphaGenome works
AlphaGenome uses a hybrid architecture combining convolutional neural networks and transformers:
- Convolutional layers detect short sequence patterns (motifs) in the input DNA
- Transformer layers propagate information across all positions, capturing long-range regulatory interactions
- Output heads convert learned representations into predictions for different functional modalities
The model was trained on thousands of experimental datasets measuring gene expression, chromatin accessibility, histone modifications, and transcription factor binding across diverse cell types. Training completed in four hours on TPUs—half the compute of the earlier Enformer model.
For variant effect prediction, AlphaGenome runs inference twice (reference sequence and alternate sequence) and computes the difference in predicted expression values.
Limitations
- Research use only: Predictions are not validated for clinical diagnostics
- Long-range interactions: Accuracy decreases for regulatory elements more than 100,000 bp from the variant
- Non-coding focus: For variants in protein-coding regions, AlphaMissense may be more appropriate
- Window size constraints: Only specific window sizes are supported (16K, 128K, 512K bp)
- Rate limits: The API is designed for small to medium analyses (thousands of predictions), not genome-wide scans requiring millions of predictions
